Lecture 11 - ghoselab.org · Outline - Nucleic Acids • Nucleotides, nucleosides, and nucleic...
Transcript of Lecture 11 - ghoselab.org · Outline - Nucleic Acids • Nucleotides, nucleosides, and nucleic...

Based on Profs. Kevin Gardner & Reza Khayat 1
Biochemistry - I
Mondays and Wednesdays 9:30-10:45 AM (MR-1307)
SPRING 2017
Lecture 11

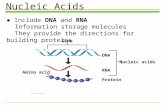
Outline - Nucleic Acids• Nucleotides, nucleosides, and nucleic acids • DNA and RNA • Properties of DNA and RNA • Structures of DNA • RNA • Hybridization • DNA sequencing (Sanger’s method) • Mutagens • Nucleotides as a source of energy and regulation
2Chapter 8: Nucleotides & Nucleic Acids

Nucleotides: Uses and Components
3
•Nucleotides: •Three components (base, phosphate, and ribose – a pentose) •Three general uses:
•Present in DNA and RNA •Act as a source of energy for living systems (ATP and GTP) •Cellular signaling, coupling extracellular stimuli with intracellular response
•Nucleosides lack the phosphate group of nucleotides
• 6 member ring • C/T/U
• The bases are divided into two parent compounds: pyrimidine and purine
8.1 Some Basics 273
Figure 8–4 gives the structures and names of the four major deoxyribonucleotides (deoxyribonu-cleoside 5!-monophosphates), the structural units of
DNAs, and the four major ribonucleotides (ribonu-cleoside 5!-monophosphates), the structural units ofRNAs.
H
C O
OH CH
CH2
2
OH
H
CO
C
CH
H
H OH
OH
OH
H
OH
OH
HH
OH
Aldehyde "-Furanose(a)
1! 1!
2!
C-2! endo
5!
4! 4!
3!
C-3! exo
C-3! endo
5!
(b) C-2! exo
Base Base
FIGURE 8–3 Conformations of ribose. (a) In solution, the straight-chain (aldehyde) and ring (!-furanose) forms of free ribose are inequilibrium. RNA contains only the ring form, !-D-ribofuranose.Deoxyribose undergoes a similar interconversion in solution, but inDNA exists solely as !-2!-deoxy-D-ribofuranose. (b) Ribofuranose
rings in nucleotides can exist in four different puckered conformations.In all cases, four of the five atoms are in a single plane. The fifth atom(C-2! or C-3!) is on either the same (endo) or the opposite (exo) side ofthe plane relative to the C-5! atom.
(a) Deoxyribonucleotides
T, dT, dTMP
Deoxythymidine
Nucleotide: Deoxyadenylate(deoxyadenosine
5!-monophosphate)
Deoxyguanylate(deoxyguanosine
5!-monophosphate)
Deoxythymidylate(deoxythymidine
5!-monophosphate)
Symbols: A, dA, dAMP
Nucleoside: Deoxyadenosine
G, dG, dGMP
Deoxyguanosine
Deoxycytidylate(deoxycytidine
5!-monophosphate)
C, dC, dCMP
Deoxycytidine
CH2O#O
OH
H
P
CH3
O#
HN
N
HH
H
H
O
O
O
OO
CH2
N
O#O
OH
H
P
NH2
O#
N
NN
HH
H
H
O
O
CH2O#O
OH
H
P
HN
H2NO#
N
NN
HH
H
H
O
O
CH2O#O
OH
H
P
NH2
O#
N
N
HH
H
H
O
O
O
(b) Ribonucleotides
O
CH2
N
O#O
OH
H
P
NH2
O#
N
NN
HH
H
O
O
CH2O#O
OH
H
P
HN
H2NO#
N
NN
HH
H
O
O
CH2O#O
OH
H
P
O#
N
N
H
H
H
O
O
CH2O#O
OH
H
P
NH2
O#
N
N
HH
H
O
O
O
OH OH OH OH
H
O
O
U, UMP C, CMP
Uridine
Nucleotide: Guanylate (guanosine5!-monophosphate)
Uridylate (uridine5!-monophosphate)
Cytidylate (cytidine5!-monophosphate)
Symbols:Nucleoside:
Adenylate (adenosine5!-monophosphate)
A, AMP
Adenosine
G, GMP
Guanosine Cytidine
FIGURE 8–4 Deoxyribonucleotides and ribonucleotides of nucleicacids. All nucleotides are shown in their free form at pH 7.0. The nu-cleotide units of DNA (a) are usually symbolized as A, G, T, and C,sometimes as dA, dG, dT, and dC; those of RNA (b) as A, G, U, and C.In their free form the deoxyribonucleotides are commonly abbreviateddAMP, dGMP, dTMP, and dCMP; the ribonucleotides, AMP, GMP,
UMP, and CMP. For each nucleotide, the more common name is fol-lowed by the complete name in parentheses. All abbreviations assumethat the phosphate group is at the 5! position. The nucleoside portionof each molecule is shaded in pink. In this and the following illus-trations, the ring carbons are not shown.
Chapter 8: Nucleotides & Nucleic Acids
• 9 member ring • A/G
• nucleotides: with phosphate • nucleosides: no phosphate
D-ribose

Ribose Conformations
4
• Conformation of ribose: four different “puckered” conformations (4 of 5 atoms in single plane), C-2’ endo or exo, C-3’ endo or exo
• Different conformations found in different DNA/RNA structures
Chapter 8: Nucleotides & Nucleic Acids

Nucleobases
5
• Weak bases • Adenine (A), Guanine (G), and Cytosine (C) are present in DNA
and RNA • Thymine (T) is only present in DNA; uracil (U) is a de-
methylated form of thymine and takes its place in RNA • Cytosine in DNA can spontaneously deaminate to uracil
Chapter 8: Nucleotides & Nucleic Acids

Nomenclature
6Chapter 8: Nucleotides & Nucleic Acids

Biophysical Properties of Nucleobases
7
• Bases (Pu/Py) have highly conjugated structures that absorb in UV ~ 260 nm • Free bases are poorly water soluble at neutral pH, more soluble in acid/base
(more charged)
What wavelengths do amino acid sidechains absorb light? Can this be usefully exploited?
Chapter 8: Nucleotides & Nucleic Acids

Chemical Modifications to Ribose & Bases
8
Similarly, the bases can be
modified.
Modification can serve to protect DNA
from digestive enzymes,
used in tRNA
6
A nucleoside’s sugar can be modified in many different ways (e.g. phosphorylation at the C5
carbon produces nucleotides)
these can be used for making DNA (nucleotides), for enzyme regulation (cAMP, last
structure)
Chapter 8: Nucleotides & Nucleic Acids

DNA and RNA: General Introduction
9
•Phosphodiester bonds link successive nucleotides in nucleic acids: the phosphate group “bridges”
the 5’-OH of one nucleotide to the 3’-OH of the next
•The phosphates have a pKa ~ 0. They always have a negative charge and form complexes with proteins and divalent cations
(e.g. Mg2+)
• DNA/RNA sequence convention is 5’ to 3’:
5’-GAATTC-3’ (Phosphate at 5’, hydroxyl at 3’ )
•DNA (deoxyribonucleic acid) lacks an OH group at C2’ •RNA (ribonucleic acid) possesses an OH group at C2’
• The ends of nucleic acids are referred to as 5’ and 3’.
This gives them directionality
Chapter 8: Nucleotides & Nucleic Acids

DNA and RNA: Stability
10
• RNA is less stable than DNA because of the ribose 2’ OH group
• Both DNA and RNA undergo slow non-enzymatic hydrolysis of the phosphodiester bond in vivo.
Chapter 8: Nucleotides & Nucleic Acids

DNA Structure: The Double Helix
11
• Primary structure = nucleotide sequence
• Secondary structure = helix
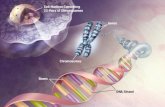
• Tertiary structure = folding in chromosomes (structure becoming understood)
• Note: (blue lines) A-T (two H-bonds) and GC (three H-bonds)
• Hydrogen bonding between specific bases is preserved (not so with RNA)
• Hydrogen bonding occurs between protonated nitrogens and carbonyls
Chapter 8: Nucleotides & Nucleic Acids

DNA structure: The double helix
12
• Structural stability due to: • Hydrophobic interactions among stacked bases pi-bonding overlap (most important) • H-bonds (less important)
• 36 Å per repeat of major turn, 3.4 Å per bp rise, 10.5 bp per turn • Schematic (a), stick (b), and space filling (c) -note the well packed structure of bases
Chapter 8: Nucleotides & Nucleic Acids

DNA structure: Torsion angles
13
• Rotation about seven different bonds (w/limited rotation about bond 4) connecting sugar (ribose) and phosphate
• Limited rotation about bond 4 gives rise to ring pucker (endo vs exo)
DNA sequence can affect these angles. This gives the DNA a particular structure that may be important for protein binding; thus gene regulation
Chapter 8: Nucleotides & Nucleic Acids

Types of DNA Double Helices
14
Three structures of DNA discovered to date (A, B and Z) A-form: believed to be a chemical artifact as a result of dehydration of B form B-form: most DNA in vivo Z-form: Found in structures near promoters (possibly responsible for gene regulation or recombination?)
• Z-DNA favored by alternating purines-pyrimidines (Pu-Py) steps
• A segment of DNA can be experimentally induced to change from B to Z conformation. When this occurs purines ”flip” from anti to syn and pyrimidines remain anti, so sugar-P backbone “zig-zags” (hence the name for Z-DNA).
Chapter 8: Nucleotides & Nucleic Acids

15
Z-form syn-purines B-form B- and Z-form
Chapter 8: Nucleotides & Nucleic Acids
B- and Z-DNA Bases

DNA Origami and Nanostructures
16
Palindromes can lead to the formation of hairpins in ssDNA or RNA). Sites recognized by many DNA binding proteins (i.e. restriction enzymes)
Cruciforms and hairpins are self complementary and have been seen in vitro, yet rarely in vivo
Two Hairpins
Chapter 8: Nucleotides & Nucleic Acids

Hoogsteen Base-pairing & Alternative Structures
17
Hoogsteen base pairs (red, right) allows for the formation of triple-helical DNA with two pyrimidine strands and one purine strand and parallel or antiparallel G-tetraplex strands.
G-tetraplex (four strands) occurs only when many guanosine bases present, including telomeres at the end of chromosomes. The dark and light blue strands are antiparallel WC base pairs and the third all-pyrimidine strand in Hoogsteen base pairs is shown in purple. Only the triplet closest to the viewer is colored.
Chapter 8: Nucleotides & Nucleic Acids

RNA: Nucleotide-based Machines of the cell
18

RNA: Nucleotide-based Machines of the Cell
19
• 3 major types of RNA (mRNA, tRNA, and rRNA); emerging importance of miRNA (micro) and many other types of ncRNAs (non-coding)
• mRNA (messenger RNA) codes for polypeptide chains: Monocistronic (prokaryotes and eukaryotes for one gene) or polycistronic (prokaryotes for multiple genes). Cistron is the genetic definition of a gene. Eukaryotic genes include mix of protein-coding segments (exons) and non-coding segments (introns).
5’ 3’5’ untranslated region
(5’ UTR)
3’ UTR
start codon (AUG) stop codon(UAA,UAG, UGA)
ORF; open reading frame
Untranslated regions serve for protein binding sites, regulation of translation, mRNA stabilization
Chapter 8: Nucleotides & Nucleic Acids
eukaryotes
prokaryotes
exonexonexonexonexon

RNA Structure
20
• Typical right-handed stacking pattern of single-stranded RNA (mRNA) •White = bases •Green = ribose and phosphate oxygen •Yellow = Pi atoms
• Secondary structure of ssRNAs: bulge, internal loop, hairpin loop
• The paired regions generally have an A-form right-handed helix (above)
“extra base”Stem loop
• ssRNA (mRNA, tRNA, rRNA) can form stem loop structures to self hybridize for greater stability
Chapter 8: Nucleotides & Nucleic Acids

tRNA: Essential for mRNA Translation
21
Phe tRNA of yeast, with unusual bp: note oxygen of a ribose phosphodiester bond in H-bonding arrangement and a ribose 2’-OH in another (red) in box
•Nearly all are L-shaped to fit into the ribosome
•AA loaded into acceptor loop (purple star)
•Anticodon loop (red star) must basepair with mRNA codon
Chapter 8: Nucleotides & Nucleic Acids

Large RNPs: Ribonuclear Protein machines
22
50S large subunit
70S prokaryotic ribosome: rRNA/rRNA / tRNA / protein
30S small subunit
translation RNA splicing – intron removal
large, well-ordered RNAs provide catalytic function & binding sites for proteins

Hybridization: How Strands Find Each Other
23
• Hybridized DNA and RNA can be denatured and renatured. The Tm depends upon the GC composition and degree of complementarity between strands. Disruption of the H-bonds is reversible.
• ssDNA to dsDNA (annealing) begins slowly by random collisions, then when in register, it “zips” up
• Central process to nucleic acid biochemistry and analytical methods
Chapter 8: Nucleotides & Nucleic Acids

DNA sequencing: Sanger method
24
• Four reactions are run, containing: — template DNA (to be sequenced) : dNTP (dATP, dGTP, dCTP, or dTTP) — oligonucleotide (short length of DNA) : DNA polymerase (enzyme) to make the DNA — nucleotide pool is spiked with radioactive di-deoxy NTP (ddNTP)
• Lack of 3’OH in ddNTP halts DNA synthesis (replication) • ssDNA strands can be separated with electrophoresis
Chapter 8: Nucleotides & Nucleic Acids

Mutagens & Mutagenesis
25
Some well-characterized non-enzymatic reactions of nucleotides: deamination and depurination. These reactions occur very slowly and lead to mutations, perhaps a link to aging and carcinogenesis. DNA repair machinery (2015 Nobel Prize in Chemistry) repair these damages
• C to U most common: (about 100/day). Corrected by DNA repair system, but not 100% fidelity!
• Methylation of Cytosine (5-Methylcytosine) is used to regulate gene transcription
• Others are less common, and corrected by the DNA repair system
• Depurination generates “apurinic” residue that is recognized by DNA repair system
• Very slow and not considered as physiologically significant
Chapter 8: Nucleotides & Nucleic Acids

Nucleotides: Use for Energy and Signaling
26
• NTPs precursors of DNA/RNA
• NTPs store energy (ATP/GTP)
• Hydrolysis of anhydride bond yields more energy than hydrolysis of the ester (below)
• ATP hydrolysis provides energy for biosynthesis
• ATP = “energy currency”
Mono, Di, and Triphosphates (NMPs, NDPs, and NTPs)
~30 kJ/mol
~14 kJ/mol
Chapter 8: Nucleotides & Nucleic Acids

Nucleotides: Use for Energy and Signaling
27
Some nucleotides are regulatory molecules: External molecules (first messengers, e.g., hormones) interact with receptors on cell surface that lead to the production of second messengers inside cell, such as cAMP and cGMP.
ppGpp is produced in bacteria in response to a slowdown in protein synthesis during amino acid starvation. This nucleotide inhibits the synthesis of the rRNA and tRNA molecules needed for protein synthesis, preventing unnecessary production of nucleic acids.
Chapter 8: Nucleotides & Nucleic Acids