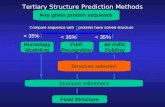
Sequence-Structure-Function Sequence Structure Function Threading Ab initio BLAST Folding:...
-
date post
21-Dec-2015 -
Category
Documents
-
view
230 -
download
0
Transcript of Sequence-Structure-Function Sequence Structure Function Threading Ab initio BLAST Folding:...
Sequence-Structure-Function
Sequence
Structure
Function
ThreadingAb initio
BLAST
Folding: impossible but for the smallest structures
Function prediction from structure – very difficult
Experimental
• Structural genomics
• Functional genomics
• Protein-protein interaction
• Metabolic pathways
• Expression data
Protein function groups• Catalysis (enzymes)• Binding – transport (active/passive)
– Protein-DNA/RNA binding (e.g. histones, transcription factors)– Protein-protein interactions (e.g. antibody-lysozyme)– Protein-fatty acid binding (e.g. apolipoproteins)– Protein – small molecules (drug interaction, structure decoding)
• Structural component (e.g. -crystallin)• Regulation• Signalling• Transcription regulation• Immune system• Motor proteins (actin/myosin)
Energy difference upon binding
Examples of protein interactions (and functional importance) include: • Protein – protein (pathway analysis); • Protein – small molecules (drug interaction, structure decoding); • Protein – peptides, DNA/RNA (function analysis)
The change in Gibb’s Free Energy of the protein-ligand binding interaction can be monitored
and expressed by the following; G = H - T x S (H=Enthalpy, S=Entropy and T=Temperature)
Protein function
• Many proteins combine functions
• Some immunoglobulin structures are thought to have more than 100 different functions (and active/binding sites)
• Alternative splicing can generate (partially) alternative structures
How to infer function
• Experiment• Deduction from sequence
– Multiple sequence alignment – conservation patterns– Homology searching
• Deduction from structure– Threading– Structure-structure comparison– Homology modelling
Metabolic Metabolic networksnetworks
Glycolysis Glycolysis and and
GluconeogenesisGluconeogenesis
Kegg database (Japan)
Gene Ontology (GO)
• Not a genome sequence database
• Developing three structured, controlled vocabularies (ontologies) to describe gene products in terms of:– biological process– cellular component– molecular function
in a species-independent manner
Gene Ontology Members
• FlyBase - database for the fruitfly Drosophila melanogaster • Berkeley Drosophila Genome Project (BDGP) - Drosophila informatics; GO database & software, Sequence Ontology development • Saccharomyces Genome Database (SGD) - database for the budding yeast Saccharomyces cerevisiae • Mouse Genome Database (MGD) & Gene Expression Database (GXD) - databases for the mouse Mus musculus • The Arabidopsis Information Resource (TAIR) - database for the brassica family plant Arabidopsis thaliana • WormBase - database for the nematode Caenorhabditis elegans • EBI GOA project : annotation of UniProt (Swiss-Prot/TrEMBL/PIR) and InterPro databases • Rat Genome Database (RGD) - database for the rat Rattus norvegicus • DictyBase - informatics resource for the slime mold Dictyostelium discoideum • GeneDB S. pombe - database for the fission yeast Schizosaccharomyces pombe (part of the Pathogen Sequencing Unit at the Wellcome Trust Sanger Institute) • GeneDB for protozoa - databases for Plasmodium falciparum, Leishmania major, Trypanosoma brucei, and several other protozoan parasites (part of the Pathogen Sequencing Unit at the Wellcome Trust Sanger Institute) • Genome Knowledge Base (GK) - a collaboration between Cold Spring Harbor Laboratory and EBI) • TIGR - The Institute for Genomic Research • Gramene - A Comparative Mapping Resource for Monocots • Compugen (with its Internet Research Engine) • The Zebrafish Information Network (ZFIN) - reference datasets and information on Danio rerio