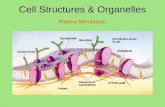
Membrane Structure and Dynamics Membrane functions - physical barrier from entry and exit form cell...
-
Upload
warren-hall -
Category
Documents
-
view
215 -
download
0
Transcript of Membrane Structure and Dynamics Membrane functions - physical barrier from entry and exit form cell...

Membrane Structure and Dynamics
Membrane functions - physical barrier from entry and exit form cell and organelles
What are membranes - Lipid bilayers with proteins imbedded or associated on either side of the membrane
Ions and polar molecules basically impermeable to membrane -
energy costs too high
Membranes - Lipid, protein and carbohydrateMembrane % Protein % Lipid % Carbohydrate
Plasma membrane 46 54 2 - 4Mitochondria 76 24 1 - 2

Membrane components - • 60 to 70% of mammalian lipids are phospholipids• Bacteria have almost no PC and are mostly PE• Neuronal tissue (myelin) PI > PC
• Alterations in lipid composition - permeability, fluidity, exocytosis, neural transmission and signaling potential
Lipid plasma membrane golgi mito nucleiP-Choline 35 45 50 62P-Ethanolamine 19 17 23 23P-Insositol 7 9 13 9P-Serine 9 4 5 4Sphingosine 18 12 3 3

• Membrane Asymmetry – P-ethanolamine and P-serine predominately faces inside of
cell– P-choline faces outside of membrane and inside of
organelles– carbohydrates of glycoproteins facing outside
• During apoptosis there is a re-arraignment of lipids where phosphatidyl serine moves to the exterior face of the membrane. One of the key signals of cell death

• Membrane Fluidity - Singer and Nickolson fluid mosaic model- allows for dynamic nature of membrane - little transition of lipids can take place without specific
enzymes to mediate transfer - flipase

• Proteins - Add function and structure to membrane• Extrinsic proteins (peripheral)
– Loosely attached to membrane– ionic bonds with polar head groups and carbohydrates– hydrophobic bonds with lipid– proteins have lipids tails– easily displaced from membrane– salt, pH, sonication

Transmembrane portion often a helix takes about 20 aa to cross membrane many proteins cross many times
odd # of transmembrane regions, why -COOH terminal usually cytosolic -NH3
+ terminal extracellular can be predicted by amino acid sequence high % of side chains will be hydrophobic Hydropathy scale used to predict
free energy change - from organic to water long regions unusual in soluble proteins• Non membrane sections often modified
lipid, carbohydrate



Intrinsic proteins - tightly bound to membrane - span both sidesProtein has both polar and hydrophobic sections removed only through
disrupting membrane with detergentsdetergents disrupt lipid bilayer and incorporate proteins and some lipids
into detergent micelles allows for purification of membrane proteins reconstitute into specific vesicles for study

Membrane associated proteinsN or C terminal modificationsTightly associates protein to membraneIsoprenylated at C Terminus
-Geranylgeranyl and farnesyl groups - from cholesterol biosynthesis
- Lovastatin inhibits post-translational modification - deterimined for Ras and pancreatic cancer.
-CAAX box - C = Cys A = aliphatic and X = various Last 4 aas are removed and new C-term is esterified with
isoprenyl
Other fatty acids can be modified at N terminus - Modification on amine or other amino acid residues- Myristoylation or Palmitoylation - usually occurs on Cys residues - highly reversible

Permeability - charged substances do not cross without help
measured by ability of small molecules to cross membranes
• Synthetic lipid vesicles formed by sonication• Measure trapped ions that cross back out into
solution• Only charged molecule that can cross easily is
water• Movement slowed by transport though two
environments• Shed layers of hydration

Summary of membrane transport
• Three types of membrane transporters enhance the movement of solutes across plant cell membranes– Channels – passive transport– Carriers – passive transport– Pumps- active transport

Channels
• Transmembrane proteins that work as selective pores– Transport through these passive
• The size of the pore determines its transport specifity
• Movement down the gradient in electrochemical potential
• Unidirectional• Very fast transport• Limited to ions and water

Channels
• Sometimes channel transport involves transient binding of the solute to the channel protein
• Channel proteins have structures called gates.– Open and close pore in response
to signals• Light
• Hormone binding
• Only potassium can diffuse either inward or outward– All others must be expelled by
active transport.

Remember the aquaporin channel protein?
• There is some diffusion of water directly across the bi-lipid membrane.
• Aquaporins: Integral membrane proteins that form water selective channels – allows water to diffuse faster– Facilitates water movement in
plants
• Alters the rate of water flow across the plant cell membrane – NOT direction

Carriers• Do not have pores that extend
completely across membrane• Substance being transported is
initially bound to a specific site on the carrier protein– Carriers are specialized to carry a
specific organic compound
• Binding of a molecule causes the carrier protein to change shape– This exposes the molecule to the
solution on the other side of the membrane
• Transport complete after dissociation of molecule and carrier protein

Carriers• Moderate speed
– Slower than in a channel
• Binding to carrier protein is like enzyme binding site action
• Can be either active or passive
• Passive action is sometimes called facilitated diffusion
• Unidirectional

Active transport• To carry out active transport:
– The membrane transporter must couple the uphill transport of a molecule with an energy releasing event
• This is called Primary active transport– Energy source can be
• The electron transport chain of mitochondria
• The electron transport chain of chloroplasts
• Absorption of light by the membrane transporter
• Such membrane transporters are called PUMPS

Primary active transport- Pumps
• Movement against the electrochemical gradient
• Unidirectional• Very slow
• Significant interaction with solute
• Direct energy expenditure

pump-mediated transport against the gradient (secondary
active transport)
• Involves the coupling of the uphill transport of a molecule with the downhill transport of another
• (A) the initial conformation allows a proton from outside to bind to pump protein
• (B) Proton binding alters the shape of the protein to allow the molecule [S] to bind

pump-mediated transport against the gradient (secondary
active transport)• (C) The binding of the
molecule [S] again alters the shape of the pump protein. This exposes the both binding sites, and the proton and molecule [S] to the inside of the cell
• (D) This release restores borh pump proteins to their original conformation and the cycle begins again

pump-mediated transport against the gradient (secondary
active transport)• Two types:
• (A) Symport:– Both substances move in the
same direction across membrane
• (B) Antiport:– Coupled transport in which the
downhill movement of a proton drives the active (uphill) movement of a molecule
– In both cases this is against the concentration gradient of the molecule (active)

pump-mediated transport against the gradient (secondary
active transport)
• The proton gradient required for secondary active transport is provided by the activity of the electrogenic pumps
• Membrane potential contributes to secondary active transport
• Passive transport with respect to H+ (proton)

The end