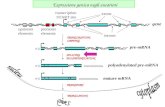
Diversité des transcrits: Les sites de polyadenylation « intergéniques »
Chapter 15 : Post-transcriptional events II: Capping and polyadenylation
description
Transcript of Chapter 15 : Post-transcriptional events II: Capping and polyadenylation

Chapter 15: Post-transcriptional events II: Capping and polyadenylation
• Cap structure


• a. a phosphohydrolysis removes the terminal phosphate from a pre-mRNA;
• b. a guanylyl transferase adds the capping GMP.
• c and d. two methyl transferase methylate the N7 of the capping guanosine and the 2’ O-CH3 group of the penultimate nucleotide.
Sequence of events in capping

• Cap structure
• DEAE-cellulose chromatographic purification of vaccinia virus cap
, - 32pGTPS-adenosyl[methyl-3H] methionine

G

• Identification of the capping substance as 7-methyl-guanosine.
• Miura and Furuichi
, - 32P-ATPS-adenosyl[methyl-3H] methionine
- 32P-ATPunable to be retained in the cap.-phosphate was Alkaline phosphatase resistant-phosphate was protected by substance X

Phosphodiesterase; phosphomonoesterase
A
Paper chromatography
electrophoresis

• Caps are made in steps : 1., a phosphohydrolysis removes the terminal phosphate from a pre-mRNA; 2. a guanylyl transferase adds the capping GMP. 3. two methyl transferase methylate the N7 of the capping guanosine and the 2’ O-CH3 group of the penultimate nucleotide.
Sequence of events in capping

Identification of ppGpC as an intermediate in reovirus cap synthesis
PPi
electrophoresis
Alkaline phosphatase, ppGpC GpC
Ion-exhanger column

Functions of Caps
Effect of cap on RNA stability -protection
Furuichi et al.
Capped - m7GpppG (green)or blocked -GpppG (blue)glycerol gradient ultracentrifugation
8 h
Wheat germ, 8 h
Remove cap or block

Effect of cap on translatabilityD. Gallie; in vivo assay

Capping of U1 snRNA is necessary for its transport to the cytoplasmHamm & Mattaj
U1- RNA Pol IIU6- RNA Pol III
U1 driven by RNA Pol III
U1-5: m2,2,7 GU6: no cap
U1- m7G (nucleus) Cytoplasm, receives other two methylation;complexed with proteins nucleus to take part in RNA splicingDoes the capping play role in transporting RNA out of the nucleus?

Mutant U1:unable to complex with proteins

• (1) protection of the mRNA from degradation; • (2) enhancement of the mRNA’s translatability; • (3) transport of the mRNA out of the nucleus;• (4) proper splicing of the pre-mRNA.
Summary- the cap provides:

Polyadenylation
• Most eukaryotic mRNAs and their precursors have a chain of AMP residues about 250 nucleotides long at their 3’ends. This poly(A) is added post-transcriptionally by poly(A) polymerase.

Sheines & Darnell
radioactively labeled HeLa cells for a short time (12 min); isolated hn RNA (nuclei) and mRNA (cytoplasm); RNase T1 (cut G), A (cut C or U) (Ap)n

Finding poly(A) at the 3’end of hnRNA and mRNA

Effect of poly(A) on translation of globin mRNA in oocytesRevel et al.
Globin mRNA(poly A+) or (poly A-) injected to frog oocytes; labeled Hb with 3H-histidine; Sephdex G-100 column filtration

Effect of poly(A) on translation of globin mRNA in oocytesRevel et al.
poly A+
poly A-

• Time course of translation of poly(A)+ and poly(A)- globin mRNA.
poly(A)+
poly(A)-

Munroe and Jacobson
Effect of poly (A) on translatability and stability of mRNAs

Effect of poly(A) on recruitment of mRNA to polysomes
Munroe & Jacobson
Poly(A) enhances lifetime and translatability. But, relative importance varies with system

• (a) cutting ,
• (b) polyadenylation,
• (c) degradation
Basic Mechanism of Polyadenylation

-Globin gene transcription extends beyond the poly(A) site.
Hofer & Darnell
Isolated nuclei from DMSO stimulated red blood cells; run-on transcription with 32P-UTP; hybridized with DNA probes (A,B,….F) of -Globin gene

Adenovirus late transcription unit
Poly(A) Poly(A)

Model 3. Transcripts are clipped and polyadenylated while transcription is still in processs
Nevins & Darnell
Model 1. Stop at the coding region and polyadenylation
Model 2. Stop at the very last end and polyadenylation

AB
CD
E
DNA probes
If model 1 is correct, then
Chance of hybridization high low
Not supported by experimental results

Nevins & Darnell

Basic Mechanism of Polyadenylation
Where?

Summary of data on 369 veterbrate polyadenylation

Importance of the AATAAA sequence to polyadenylation
Fitzgerald & Shenk
Recombinant SV40 virus
But, AATAAA is not sufficient. Deletion of immediate down stream region of the site can disrupt the polyadenylation

AATAAAA-N(23/24)-GT rich region-T rich region
Gil and Proudfoot
-globin gene


Splicing happens before polyadenylation

Cleavage and poly-adenylation of a pre-mRNA
A model for the pre-cleavage complex

Both PAP and CPSF are necessary for polyadenylation
M. Wickens et al.
Initiation of Polyadenylation

Polyadenylation has two phases Sheets & Wickens

CPSF binds to the AAUAAA motif
Keller et al.
35 and 160 Kd proteins

• Polyadenylation requires both cleavage of the pre-mRNA and polyadenylation at the cleavage site. Cleavage in mammals requires : CPSF, CstF, CF1 and CFII, and poly(A) polymerase (PAP).
• Polyadenylation has two phases. Once the poly(A) reaches about 10 nt in length, further polyadenylation becomes independent of the AAUAAA signal and depends on the poly (A) itself.
Summary

Purification of poly(A)-binding protein (PABII)E. Wahle
Elongation of Polyadenylation
49 Kd proteinActivity assay
Nuclear protein

Effect of CPSF and PABII on polyadenylation

• Elongation of poly(A) requires PAB II. This protein binds to a pre-initiated oligo (A) and aids poly(A) polymerase in elongating poly(A) up to 250 nt.
• PAB II acts independently of the AAUAAA motif. It depends on poly(A), but its activity is enhanced by CPSF.
PAPCPSF
PABII
CFI, II, CstF Summary


Architecture of PAP
Specific polyadenylation carried out by full-length andC-terminally truncated PAP
Manley et al.

Shortening of cytoplasmic poly(A)
Sheines & Darnell Turnover of Poly(A)
48 h labeling
Cytoplasmic poly(A) RNA
Nuclear poly(A) RNA
• Summary - Poly(A) turns over in the cytoplasm. RNase tears it down, and PAP builds it back up. When the poly(A) is gone, the mRNA is slated for destruction.
12 min labeling

Dependence of PAN on PAB I,and distributive nature of PAN
Sachs et al.

Biphasic de-adenylation
Sachs et al.

• Cytoplasmic deadenylation is carried out by PAN (poly(A) nuclease), in conjunction with PAB I (poly(A) binding protein).
• This reaction is biphasic. Rapid and slow phases (terminal 12-25 nt).
Summary-

Various rates of de-adenylation in yeast mRNAs
Sachs et al.

A sequence in mRNA 3’UTR that inhibitsterminal deadenylation
Summary of 3’UTR mutations and theireffects on de-adenylation
Sachs et al.

• Deadenylation is not equally efficient for all mRNAs.• The 3’UTR controls the efficiency of de-adenylation. An adenin
e-uridine-rich (ARE3) about 60 nt upstream of the poly(A) tail is a sensitive site.
Summary-

Cytoplasmic poly-adenylation
Maturation-specific poly-adenylation in frog oocytes; Maternal RNA[Poly(A)-]; D7 RNA polyadenylated.

Cytoplasmic poly-adenylation
Maturation-specific poly-adenylation of two RNAs
Wickens et al.

UUUUUAU confers maturation specific poly-adenylation
Abolition of maturation specificpoly-adenylation by mutations in the AAUAAA motif

The Effect of the Cap and Poly(A) on Splicing
• (1) protection of the mRNA from degradation;
• (2) enhancement of the mRNA’s translatability;
• (3) transport of the mRNA out of the nucleus;
• (4) proper splicing of the pre-mRNA.
The Cap function:

Shimura et al
Production of capped and uncapped splicing substrates
Effect of cap on splicing a substrates with two introns
HeLa nuclear extract

Effect of CBC on splicing and pre-splicesome formation
Mattaj et alCap binding complex: CBP80 and CBP20
Activity assayWestern blotting
Splicesome complex assay

• Summary- Removal of the first intron from model pre-mRNAs in vitro is dependent on the cap. This effect may be mediated by CBC that is involved in spliceosome formation.

Effect of polyadenylation on splicing a pre-mRNA with a single intron.Niwa & Berget

Niwa & Berget
Effect of poly-adenylaton on splicing a two-intron substrates

• Summary- polyadenylation of model substrates in vitro is required for active removal of the intron closest to the poly(A). However, splicing any other introns out of the these substrates occurs at a normal rate even without polyadenylation.