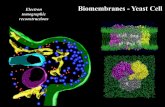
yeast cell
description
Transcript of yeast cell

yeast cell
What is a cell?
material and
energy
cell wall (protection)
cell membrane (import + export, signaling)
cytoplasm (energy production, growth)
nucleus (heredity)

What cells can do?
move
Shape change
A
B
CE
DFeeding + excreting
signaling
Y

The information flow in cells (central dogma)
DNA
DNA
DNA
DNADNA
DNA replication
+
DNA
DNA
RNA
RNA
DNA
Transcription
genetic code
protein

DNA replication
cell
division
chromosomesegregation
xeroxmachine
DNA
collatingmachine

The central dogma
S P
gene
transcript
protein
metabolism

genome
transc
ripto
me
prote
ome
metabolome
Beyond the central dogma
S P
networks

G1 G2

SPF MPFCdk
CycBS
Cdk
CycBM

SPF
MPFCdk
CycBS
Cdk
CycBM
G1 G2
Start
G2/M
exit

cell
division
controlbox
DNA replication
chromosomesegregation

ubiquitindependentproteolysis
reversibleTyrosinephosphorylation
binding of astoichiometricinhibitor

SPF
MPF
cytoplasmic mass doubling time
Cdk
CycBS
Cdk
CycBM
G1 G2
Start
G2/M
exit

0
50
100
150
350
400
WT wee1 - cki -
+ -+ + -
cycl
e t
ime (
min
)
cki –
wee1 -
G1
S
G2
M
cdc25 -
G1
S
G2
M
G1S
G2
M
G1
S
G2
M
G1
S
G2
viability
cytoplasmic mass doubling
time
Balanced growth and division

0
50
100
150
350
400
0
50
100
150
350
400
WT wee1 - cki -
+ -+ + -
cki –
wee1 -
G1
S
G2
M
cdc25 -
G1
S
G2
M
G1S
G2
M
G1
S
G2
M
G1
S
G2
cell
volu
me a
t div
isio
n (
fl)
Balanced growth and divisioncy
cle t
ime (
min
)
viability

G1 G2
SPF
pheromone
-
Cdk
CycBS
Cdk
CycBM
MPF

G1 G2
inhibitor-
SPFCdk
CycBS
Cdk
CycBMStart
MPF

G1 G2
inhibitor-
Start
G2/M
SPFCdk
CycBS
MPFCdk
CycBM


Cdk
CycB
G1 G2
cell mass doubling time
+ +Sta
rt
G2/M
exit

Sta
rtCK
I
Cdk
CycB
CK
I
P
CK
I
AA
Exit
APCM
Cdk
G2/M
P
Cdc25
Wee1Cdk
CycB
APCG1
Cdk
CycB
AA
Cdk
CycB
inactive
Wee1P
inactive
Cdc25
P
inactive
APCM
delay
inactive
APCG1inactive

?
fission yeast budding yeast

Mutants in Fission Yeastgene viability trait
cdc2- no block in G1 and G2cdc13- no endoreplicationrum1- yes sterileste9- yes sterilewee1- yes small cellscdc25- no block in G2
cdc2op yes wtcdc13op yes wtrum1op no endoreplicationste9op no endoreplicationwee1op yes large cellscdc25op yes small cells
wee1- rum1 no extremely smallwee1- cdc25 yes quantized cycleswee1- cdc25op no mitotic catastrophe

0 50 100 1500.0
0.5
1.0
0.0
0.2
0
1
0
1
1
2
period of oscillation= cycle time= mass doubling time (tD)
dm/dt = m where = ln2/tD
rate ~ mass
APCM
Cdk
CycB
APCG1
CK
I
S/G2G1M
time (min)
M
CycB
periodtD
mass (m)
tim
e

0 50 100 1500.0
0.5
1.0
0.0
0.2
0
1
0
1
1
2
period
dm/dt = m where = ln2/tD
APCG1
Cdk
CycB
APCM
CycB
CK
I
S/G2G1M
time (min)
M
mass
Size control over the cell cycleSize control over the cell cycle
rate ~ mass
periodtD
mass (m)
tim
e

Inside view ?

Sta
rtCK
I
Cdk
CycB
CK
I
P
CK
I
AA
Exit
APCM
CdkP
Cdc25
Wee1Cdk
CycB
APCG1
Cdk
CycB
AA
Cdk
CycB
Wee1P
Cdc25
P
APCG1
APCM
delay
cell mass
Cdk/C
ycB
cell mass
Cdk/C
ycB
cell mass
Cdk/C
ycB
G2/M

0 1 2
0.0
0.2
0.4
1.4
1.6 stable s.s.unstable s.s.min/max
Cd
k/C
ycB
G1
S/G2
M
cell mass
Bifurcation diagram of the cell cycle control networkBifurcation diagram of the cell cycle control networkBifurcation diagram of the cell cycle control networkBifurcation diagram of the cell cycle control network

0 1 2
0.0
0.2
0.4
1.4
1.6 stable s.s.unstable s.s.min/max
Cd
k/C
ycB
G1
S/G2
M
cell mass
Cell cycle regulation on the bifurcation diagram Cell cycle regulation on the bifurcation diagram Cell cycle regulation on the bifurcation diagram Cell cycle regulation on the bifurcation diagram
dividingcellnewborn
cellS
tart
G2/M
Exit

G1S/G2
M
0
0.4
0 2 431 5
G1 checkpoint
cell mass
Cdk/C
ycB
G1S/G2
0.4
0
0.8
1.2
1.6
0 2 431 5
M
metaphase checkpoint
cell mass
Cdk/C
ycB
G2
/M
Exit
G1S/G2
M
0
0.4
G2 checkpoint
cell mass
Cdk/C
ycB
0 2 431 5