Toc33G Toc159G Toc120A Toc132A - Plant Physiology · 2010. 5. 10. · Toc33G Toc159A Toc132A...
Transcript of Toc33G Toc159G Toc120A Toc132A - Plant Physiology · 2010. 5. 10. · Toc33G Toc159A Toc132A...

Toc33G
Toc159A
Toc132A
Toc120A
Toc159G
Figure. S1. Prediction of CK2 phosphorylation sites. NetPhosK 1.0 has been used to predict CK2 phosphorylation sites for the A-domains of Arabidopis Toc120, Toc132, and Toc159, and the G-domains of Toc33 and Toc159. Serine phosphoryla-tion sites are indicated in red and threonine phosphorylation sites in gray. Sequences used for the in silico prediction are identical to the sequences of the construction used in the in vitro phosphorylation assay, see Figure 7 B.
1- -265
727- -1093
1- -343
1- -740
1- -431

Toc159 full lengthA-dom fragment
2001169766
45
31
proto
plasts
P1 1.50
0xg
SN1 1.50
0xg
116
116976645
31
kDa
BSA (breakage buffer)LSU
CAB
1 2 3 4 5 6
IVTIVT -TL +TL+TL-TL
2001169766
45
31
kDa kDa
A
B
protoplastsrupturing device
broken protoplasts1.500 x g, 4 min
P1 SN1
Toc15GM
Toc159M
G MA G M1 1503 728 1503
Toc159
Toc15GM
Toc159M
1 2 3
Toc159 Toc15GM
C
anti-Toc159A
Phosphor-imager
Amido Black
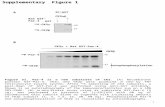
Supplemental Figure S2. Toc159 cleavage. (A) A Toc159 processing activity in isolated chlo-roplasts. [35S]-methionin-labelled, in vitro translated (IVT) Toc159 or Toc159GM were incubated with isolated chloroplasts. Chloroplasts were re-isolated and incubated with (+TL) or without (-TL) thermolysin. Samples were analyzed by SDS-PAGE and PhosphorImaging. (B) Occurrence of the A-domain fragment after gentle cell disruption. Protoplasts of wild-type Arabidopsis plants were disrupted with a rupturing device employing 23 and 18 µm nylon mesh in two layers (Smith et al., 2002). Organelles were separated by centrifugation. 50 µg of protoplast, pellet and superna-tant protein were analyzed by Western Blotting with anti-Toc159A. (C) An annotated semi-tryptic peptide identified in the SN100 soluble fraction. To identify the most C-terminal amino acid of the soluble A-domain, MS/MS spectra were searched by Mascot for peptides that show tryptic specifi-city at one terminus only, and where the other terminus may be the result of non-tryptic cleavage. The semi-tryptic peptide mapping to amino acids 733-744 of atToc159 was identified in the SN100 fraction of three independent purifications with ion scores of 37, 42 and 47.

locus peptide sequence title query number AT4G02510 VGADDLpSDSEK F059194 q801_p1 AT4G02510 VVEGDpS AEEDENKLPVEDIVSSR F075163 q1354_p1 AT4G02510 KVVEGDpSAEEDENKLPVEDIVSSR F075163 q1360_p1 AT4G02510 FDQIGDDDpSGEFEPVSDK F059194 q1585_p1 AT4G02510 VGADDLSDSEKEKPNLVGDGK F050642 q1602_p1 AT4G02510 ASSGIEAHSDEANISNNMSDR F059194 q1640_p1 AT4G02510 ASSGIEAHSDEAN ISNNMSDR F059194 q1641_p1 AT4G02510 VVEGDpSAEEDENKLPVEDIVSSR F059194 q1672_p1
Supplemental Table S1: List of phosphorylated peptides presented in the main text (Tab.I) and the respective query number as its spectra identifier.
Supplemental Spectra S1: Pages 3 to 10Spectra of phosphorylated peptides as listed before. Unambiguous phosphorylation sites are assigned by " p " prior to the phosphorylated aminoacid. The p-site assign-ment " * " in the peptide sequence on each spectra page was automatically annotated without validation.

m/z
Rel
ativ
e In
tens
ity
228 406 584 761 939 1117
0
20
40
60
80
100
0
6156
12312
18468
24624
30780
● ●●
● ●
●
●
● ●●
●
●
●●
●
●●
●
●●● ●
●● ●
●
●● ●
●●
●
●●
●
●●
●●
●
●●
●
●
●
●● ●
●
●
●●
●
●
y(3)
y*−98(4)
y(6)
y*−98(7)
y−98(7)
y−98(8)
y(8)y−98(6)
y(7)
y*−98(8)
y(5)
y*(11)++
y_0(11)++
y*−98(11)++
y_0−98(11)++
y−98(11)++
363.
4570
936
3.45
709 75
8.63
591
758.
6359
1
775.
7818
6
890.
8259
4
988.
7398
660.
6883
7
873.
8252
887
3.82
528
645.
6223
599.
5510
859
9.55
108
550.
803
550.
803
559.
7746
6
a(6)
b_0(6)
b(10)
a_0(6) b−98(10)
a(10)++
b_0(5)
b−98(8)
b(8)b(6)
b(4)
b(5)
b*(11)++
b_0(11)++
b(11)++
b−98(11)++
543.
5436
5
553.
6396
3
1069
.971
2
525.
4536
8
971.
9331
6
521.
4496
4
440.
3908
8
755.
7938
4
853.
7042
6
571.
5636
9
343.
4491
6 458.
5234
1
590.
5979
359
0.59
793
599.
5510
8
550.
803
title: F059194query number: q832_p1mascot Score: 85.73precursormass/charge: 608.242810,2+delta score; pep rank1 − pep rank2: 33.9peptide sequence: VGADDL*SDSEK
y−98(5) b−98(7)y(5)
●●
●
[M+2H]−98

m/z
Rel
ativ
e In
tens
ity
338 590 841 1093 1344 1596
0
20
40
60
80
100
0
1219.6
2439.2
3658.8
4878.4
6098
●●
●
●
●
●●
●
●
●●
●
●
●●
●
●
●●
●
●● ●
●
●
●
●
●
●
● ●
●
●
●
●
●
●
●●
●
●
●
●
●
●
●
●
●
●●
●
●●
●
●
● ●●
●
●
●
● ● ●
●
●
●
●
●
●
●
●
●
●
●
● ●
●
●
●
●
●
●
●
●
●
●
●
●
●
● ●
●
●
●
●
●
●
●
●
●
●
●
y_0(13)++
y*−98(21)++
y_0−98(21)++y*−98(8)
y(21)++y−98(22)++
y−98(21)++y(7)
y(20)++
y(9)++
y(14)++
y*(21)++
y*−98(22)++
y_0(21)++y_0−98(22)++
y(9)734.
6896
5
1141
.867
511
41.8
675
789.
6618
4
1199
.466
311
99.4
663
1150
.874
5
805.
5226
8
1135
.231
4
501.
5168
800.
2479
2
1190
.854
411
90.8
544
1190
.854
411
90.8
544
1001
.638
8
b_0−98(11)
b−98(8)
b(19)++b−98(20)++
b_0(20)++
a*(15)++
a_0(15)++
b−98(11)
b*(14)++
b*−98(15)++
b_0(14)++
b(14)
b−98(14)++
a(14)++
b(14)++
b−98(15)++
1124
.984
9
769.
4314
1
1075
.264
210
75.2
642
1114
.699
6
824.
3531
282
4.35
312
1141
.867
5
789.
6618
478
9.66
184
789.
6618
4
1595
.677
2
749.
2216
8
784.
6995
7
798.
5633
279
8.56
332
title: F075163query number: q1354_p1mascot Score: 76.76precursormass/charge: 866.061710,3+delta score; pep rank1 − pep rank2: 63.39peptide sequence: VVEGD*SAEEDENKLPVEDIVSSR
y(18)++●
[M+3H]−98

m/z
Rel
ativ
e In
tens
ity
309 589 868 1147 1426 1706
0
20
40
60
80
100
0
8384
16768
25152
33536
41920
●●
●
●●
●
● ● ●●
●
●● ● ●
●
●
●●
●
●
●
●
● ●
●●
●
●
●
●
● ●●
●
●
●●
●
●
●
●
●
●●
●
●
●
●
●●●●
●● ●
● ● ●
●
● ●
●
●● ●
● ●
●
●
●●
●
●
●●
● ●●
● ●
●
●●
●
●
●●
●
●
●
●
●●
●● ●
●
●●
●
●●
●
●
●●
●
●
●●
●●
●
●●●
●
●
●
●●
y(23)++
y(5)
y*−98(6)
y_0(22)++y(17)++
y(18)++
y*−98(19)++
y_0−98(19)++
y(21)++
y−98(22)++
y(20)++
y(22)++
y−98(23)++
y(7)
y(10)
y(9)
1299
.015
5
561.
4644
356
1.46
443
1239
.524
8
965.
5858 10
49.0
353
1049
.035
310
49.0
353
1199
.910
811
99.9
108
1135
.398
8
1249
.234
1249
.234
805.
2995
7
1114
.714
3
1001
.576
6
a(15)++
b*(8)
b_0(8)
b(6)
b_0−98(15)++b(12)
b_0−98(11)++
b(11)
b(14)++
b*−98(15)++
b*(15)++
b*−98(16)++
b_0(15)++
b−98(15)++
b(15)++
a_0(18)++
848.
7045
484
8.70
454
848.
7045
4
628.
4139
2
804.
4290
8
1368
.533
1
561.
4644
3
1239
.524
8
805.
2995
780
5.29
957
853.
7804
185
3.78
041
853.
7804
1
813.
7185
3
862.
7162
8
1001
.576
6
title: F075163query number: q1360_p1mascot Score: 89.63precursormass/charge: 908.759280,3+delta score; pep rank1 − pep rank2: 72.73peptide sequence: KVVEGD*SAEEDENKLPVEDIVSSR
y(18)++
b−98(7)++ b−98(7)●
● ●
[M+3H]−98

m/z
Rel
ativ
e In
tens
ity
350 623 896 1168 1441 1714
0
20
40
60
80
100
0
385
770
1155
1540
1925
● ●●
● ●
●
●
●
●●
●●
●
●
●●
●
●
●
● ●
●
●
● ●
●
●
●
● ● ●
●
●
●
● ●
●●
●
●
●
●
●
● ●
●
●
●
●
●
●
● ●● ● ●
●
●
●
●
●
●
●
●
y−98(9)
y*(11)
y(9)++
y*(16)++y_0(16)++
y(9)
y*(6)
y_0(6)
y(6)
y(12)
y*−98(13)
y(7)
y*−98(18)++
y_0−98(18)++
y(5)
y−98(18)++
910.
0043
8
1272
.522
9
504.
6275
5
900.
8336
890
0.83
368
1008
.044
5
656.
7902
765
6.79
027
674.
7612
3
1404
.415
714
04.4
157
821.
9768
7
982.
9261
498
2.92
614
545.
6616
6
992.
0103
8
b(3)
b(11)
b*(16)++
b_0(16)++
a_0(15)++
a_0(18)++
b(8)b(7)
b*−98(8)
b_0−98(12)
b*−98(13)
b_0−98(13)
b−98(12)
b_0−98(7)
b*(13)b−98(18)++
391.
4979 12
59.9
282
900.
8336
890
0.83
368
843.
0711
5
1008
.044
5
906.
3201
8
791.
7043
279
1.70
432
1290
.120
8
1420
.058
1420
.058
1309
.044
3
674.
7612
3
1518
.219
6
982.
9261
4
title: F059194query number: q1585_p1mascot Score: 37.48precursormass/charge: 1040.407300,2+delta score; pep rank1 − pep rank2: 23.68peptide sequence: FDQIGDDD*SGEFEPVSDK
y−98(10)●
[M+2H]−98

m/z
Rel
ativ
e In
tens
ity
228 501 774 1046 1319 1592
0
20
40
60
80
100
0
26940
53880
80820
107760
134700
●
●● ●● ●● ●●
●● ●●
●● ●
●
●●
●
●
●
●●● ● ● ● ●●
●
● ●
●
●●
●●
● ● ●● ●
● ●●
●
●
● ●●●
●● ● ●
●
●● ●● ●●
●●
●● ● ●● ●●
●
●
●
●
●●
●
● ●● ● ● ●
●
●
●●● ● ● ●
●●
● ● ● ●● ●
● ●●●
●●
● ● ●●
● ●● ●● ●
●
●●
●
●
●
●●
●
● ● ● ●●
●●
●● ●● ● ●●
●
● ●
● ● ●● ●● ●
y−98(8)
y−98(15)++
y*−98(15)++
y_0−98(15)++
y−98(19)++
y(16)++
y*−98(17)++
y_0−98(17)++
y(17)++
y*−98(18)++
y−98(20)++
y−98(18)++
y−98(17)++
y(8)
y−98(16)++
y_0−98(14)++
701.
2635
5
793.
2822
378
4.42
358
784.
4235
8
999.
9456
6
898.
5214
889
8.52
148
898.
5214
8
956.
2191
495
6.21
914
1028
.897
9
964.
7992
6
907.
3411
2
799.
5572
9
849.
8199
7
700.
0958
9
b_0−98(13)++
b(16)++
b−98(17)++
b*(19)++
b_0(19)++b(18)++
b*−98(19)++
b_0−98(19)++
b−98(9)
b−98(15)++
b(13)++b−98(13)++b(19)++
b−98(19)++a(13)++b*(13)++
669.
6380
1
889.
9852
988
9.98
529
1016
.802
810
16.8
028
967.
7959
896
7.79
598
967.
7959
8
841.
9421
4
784.
4235
8
728.
1546
8
679.
1576
9
1025
.826
8
976.
6219
1
713.
5780
371
9.52
607
title: F050642query number: q1602_p1mascot Score: 68.72precursormass/charge: 752.015930,3+delta score; pep rank1 − pep rank2: 6.05peptide sequence: VGADDL*SDSEKEKPNLVGDGK
b−98(7)y−98(15)++y(15)++
●● ●
[M+3H]−98

m/z
Rel
ativ
e In
tens
ity
290 525 760 994 1229 1463
0
20
40
60
80
100
0
12766
25532
38298
51064
63830
●●
●
●
● ●
●
●
●
● ●●●
●
●●
●
●
●●
●
●
●
●
●
●
●●
●
●
● ●
●
●
●
●
●
●
●● ●
●● ●
●
●
●
●
●●
●
●●
●
●
●
●
●
●
●
●
●
●
●
●
●
●
● ●
●
●
●
● ●●
●
●●
●
●
y*(10)++
y(7)++
y(4)
y_0(9)++
y−98(7)
y(8)++
y*(16)++y*(8)
y(10)++
y(5)y*(12)++
y(8)
y*(6)y(6)
y_0−98(7) y(7)
560.
8954
5
420.
6601
3
524.
6162
852
4.61
628
740.
8134
9
477.
1507
5
935.
1783
293
5.17
832
569.
5786
8
638.
7220
9
683.
1845
6
952.
9550
9
735.
5824
8
752.
7161
4
723.
4491
1
839.
7779
6
b−98(12)++
b(12)++ b(20)++
b*(13)++b_0(13)++
b−98(14)++
a*(14)++a(14)++
b*−98(15)++
b_0(8)a*(15)++
a_0(15)++
b(13)++
b*(14)++
b−98(17)++b(14)++
569.
5786
8
618.
3541
1
1064
.146
666.
5826
666.
5826
683.
1845
6
709.
6528
3
718.
1283
718.
1283
735.
5824
8
752.
7161
475
2.71
614
675.
6338
6
723.
4491
1 839.
7779
6
732.
3936
9
title: F059194query number: q1640_p1mascot Score: 49.02precursormass/charge: 767.640560,3+delta score; pep rank1 − pep rank2: 7.88peptide sequence: ASSGIEAH*SDEANISNN*MSDR
b−98(9)++●
[M+3H]−98

m/z
Rel
ativ
e In
tens
ity
326 554 781 1009 1236 1463
0
20
40
60
80
100
0
9346
18692
28038
37384
46730
●
●
●
●●
●
●
●
●
● ●
●
●
●
●
●●
●
●
●
●
●
●
●
●
● ●
● ●
●●
●
●
●
● ● ●
●
●
●●
●
● ●●
●
●●
●●
● ●
●
●
●
●●●
●
●
● ●
●●
●
●
●
●
●
●
●
●●
●
●
●
●
●
●
●●
●●●
●
●
●
●
●
●
●
●
●
●
●
●
y*(10)++
y*(16)++
y(3) y(8)++
y(9)
y−98(20)++
y*(9)++
y_0(9)++
y*(12)++
y*(7)y*(6)
y(5)
y(8)
y(6)
y_0−98(7) y(7)
560.
8504
2
895.
1857
5
377.
4264
4
477.
1096
6
1066
.985
410
66.9
85452
4.85
287
524.
8528
7
683.
0506
4
822.
7598
735.
6313
8
638.
6735
4
953.
0109
5
752.
7654
8
723.
5759
3
839.
7541
5
b_0−98(11)++
b(12)++
b*−98(13)++
b_0−98(13)++
b−98(14)++
b−98(9)
b_0(16)++
b−98(8)
a*(14)++a_0(14)++
b(13)++
a(14)++
b*−98(15)++
a*(15)++
a_0(15)++
b*(14)++
524.
8528
7
617.
8261
361
7.82
613
617.
8261
3
683.
0506
4
822.
7598
822.
7598
735.
6313
8
709.
1449
709.
1449
675.
8296
7 718.
2354
471
8.23
544
752.
7654
875
2.76
548
723.
5759
3
title: F059194query number: q1641_p1mascot Score: 52.90precursormass/charge: 767.642180,3+delta score; pep rank1 − pep rank2: 0peptide sequence: AS*SGIEAHSDEANISNN*MSDR
b(3)●
[M+3H]−98

m/z
Rel
ativ
e In
tens
ity
328 582 835 1089 1343 1596
0
20
40
60
80
100
0
656.4
1312.8
1969.2
2625.6
3282
●
●●
●
●●
●
●
●
●
●
●
●
●
●●
●
●
●
●
●
●
●●
●
●
●
●
●
●●
●
●
●
●
●
●
●
●●
●
●
●
● ● ●●
●
●
●●
●
●●
●
●
● ●●
●●
●
●
y*−98(6)
y(11)++
y(4)
y(18)++
y*−98(19)++
y(16)++
y(6)
y(7)
y*−98(8)
y−98(21)++
y(9)++
y(14)++
y*(21)++
y*−98(22)++
y_0−98(22)++
y(9)
561.
7605
622.
2318
4
448.
5624 10
49.5
244
1049
.524
4
930.
3890
7
676.
8874
805.
8002
5
789.
9749
1151
.020
2
501.
7309
9
800.
9749
4
1191
.441
411
91.4
414
1191
.441
4
1002
.066
6
b−98(23)++b(12)
b−98(11)
b*(20)++b_0−98(7)
b(13)++b−98(20)++
b*(14)b*−98(15)
b_0(14)
b(14)
b(20)++
b_0−98(11)b*(14)++b−98(14)++ b−98(16)++ 12
40.8
145
1355
.033
3
1142
.735
6
1116
.339
5
622.
2318
4
742.
2049
3
1075
.606
5
1578
.218
615
78.2
186
1578
.218
6
1596
.240
3
1124
.960
211
24.9
602
789.
9749
749.
8348
1
847.
5266
2
title: F059194query number: q1672_p1mascot Score: 36.89precursormass/charge: 866.059720,3+delta score; pep rank1 − pep rank2: 25.44peptide sequence: VVEGD*SAEEDENKLPVEDIVSSR
y(18)++●
[M+3H]−98