Supplementary Information Structure of the VP16 ......Supplementary Information Structure of the...
Transcript of Supplementary Information Structure of the VP16 ......Supplementary Information Structure of the...

Supplementary Information
Structure of the VP16 Transactivator Target in ARC/Mediator
Alexander G Milbradt1, Madhura Kulkarni2,3*, Tingfang Yi1*, Koh Takeuchi1,4, Zhen-Yu J Sun1,
Rafael E Luna1, Philipp Selenko1,4, Anders M Näär 2,3 and Gerhard Wagner1
1Department of Biological Chemistry and Molecular Pharmacology, Harvard Medical School,
240 Longwood Avenue, Boston, MA 02115, 2Massachusetts General Hospital Cancer Center,
Building 149, 13th Street, Charlestown, MA 02129, 3Department of Cell Biology, Harvard Medical
School, 240 Longwood Avenue, Boston, MA 02115.
4Present addresses: Biomedical Information Research Center, National Institute of Advanced
Industrial Science and Technology, Aomi 2-3-26, Koto, Tokyo, 135-0064 Japan
[email protected], T.K.; Leibniz Institute of Molecular Pharmacology (FMP) Department
of NMR-assisted Structural Biology, Robert-Roessle-Strasse 10 13125 Berlin, Germany
[email protected], P.S.
* These authors contributed equally to this work.
Correspondence should be addressed to A.M.N. ([email protected]) or G.W. ([email protected]).
Nature Structural & Molecular Biology doi:10.1038/nsmb.1999

a
bCore domain TADVP16 1 490
MED25 NTD VBD391 553
1 747
411
420 430 440 450 460
HSV1 TAP TDVSLGDEL L GE V M DALDDFDL MLGD P PG T HD YS P H D D A AHA D GDS G F P SAP Chimpanzee_sv TAP TDVSLGDEL L GE V M DALDDFDL MLGD P PG T HD YS L R G E D TPV D VEF S M . PVP HSV2 TAP TDVSLGDEL L GE V M DALDDFDL MLGD P PG T HD YT I R D E D TPA E VES S M . PVS
470 480 490
HSV1 GALD DFEFEQMFTDALGID GG MA EYChimpanzee_sv GALD DFEFEQMFTDALGID GG VA DFHSV2 GALD DFEFEQMFTDALGID GG VD DF
MED25 NK LAWSGV EWQE P R LP YVN GE L T QWPGQQSVS L L KPK ASVDANTKLT S CQV H N K EPTOV1_NTD NK LAWSGV EWQE P R LP YVN GE L T QWP.EHRLS L L KRR YS.DSTAKLK T CQA Q N E DPTOV1_CTD NK LAWSGV EWQE P R LP YVN GE L T QWP..QIVN F M .PR ...EPNSRSK W SHV Q I R E
MED25 KL MQLIPQQLLTTL PLFRNS QFHFT D LK L RIM NGFAGC FQ I G RMV NK LES G Y G VH PTOV1_NTD KL MQLIPQQLLTTL PLFRNS QFHFT D LK L RIM NGFAGC FQ I G QLA NR CDS G C G ML PTOV1_CTD KL MQLIPQQLLTTL PLFRNS QFHFT D LK L RIM NGFAGC FR Y V RLV .K LET S C D VH
400 410 420 430 440
450 460 470 480 490 500
510 520 530 540 550
MED25 CE RVLMLLYSS KKIF GLIP DQ FV IR VI QPHTAP V K M Y SG NG Q TNHK VQQQKLEPTOV1_NTD CE RVLMLLYSS KKIF GLIP DQ FV IR VI QPHISP V K M Y SG SA Q TTRK A......PTOV1_CTD CE RVLMLLYSS KKIF GLIP DQ FV IR VI QSYKAS I E I H GN NG R ANQQ VLQRNLE
β1 β2
β3 α1 β4 α2
β6 α3
β5
β7
TADcTADn
TADc
Supplementary Figure 1. Sequence alignments of the MED25 VBD and VP16 TAD. (a) Sequence of the human MED25 VBD aligned with the homologous C- and N-terminal domain of human PTOV1. Secondary structure elements are shown above the sequences. (b) Sequence of the VP16 TAD from Herpes simplex virus I aligned with sequences of TADs from other viruses. Alignments were visualized with ESPript1. Conserved residues are highlighted in red.
Nature Structural & Molecular Biology doi:10.1038/nsmb.1999

E 442
N467
A504
R 538
M470
H544
H435
V 508
K 398
S 416
G462
V 417
C 429
E 410
L513
K 411
T 424Q409
V 534
L448
R 488L461
S 396
T 476
Y 487
99
N477
N535
F533
E 553
Q548
V 431N438
W 402
K 545
V 395
204
S 482
A415
Q530
K 518
W NE 402
V 405F500
G524
202
Q455
T 441
S 468
F522
Q456
Q392
Y 432F473
I541
V 510R 469
V 47181
T 503
S 531
G436
S 394
I449
M450
Q430
V 540
R 425
T 421
W NE 444
R 509I537
E 507
K 447
T 542
F494
160
L457
K 440
58
C 497
N492
V 547
73
I453
L464Q446
L552Q451
K 484M490
I521
208
L525
Q546 107
Q443
L400
M523
T 459
S 517
V 433
G532
L514
V 498
K 520
A419W 408
A495
L406
199
163 L423 K 422
S 426
L439
N434
E 437
D418
N397
S 403
D479
L511
G491
G496
W NE 408
K 519
M512
G391
S 516
Q539
T 460
F465
L399
L486
Q393
K 551L480
E 407
K 478
F390
E 481
L427
G493
H499
L458
H474 Q472
I489
Y 528
G404
K 413
L483
N420
G485
N543
I526
D529
W 444Y 515
G536
A401
193
L452
R 466
F475
10.5 10.0 9.5 9.0 8.5 8.0 7.5 7.0 6.5
105
108
111
114
117
120
123
126
129
132
15N
(pp
m)
H (ppm)1
Supplementary Figure 2. 1H-15N-HSQC spectrum of the MED25 VBD with the assigned residue numbers.
Nature Structural & Molecular Biology doi:10.1038/nsmb.1999

Loop β2-β3435 - 442
Loop β5-β6500 - 508
0
20
40
60
80
140
160
180
200
391
394
397
400
403
407
410
413
416
419
422
425
428
431
434
437
440
443
447
450
453
456
459
462
466
469
472
475
478
481
484
487
490
493
496
499
502
507
510
513
516
519
522
525
528
531
534
537
540
543
546
549
552
a
b
Loop β2-β3435 - 442
Loop β2-β3435 - 442
Loop β5-β6500 - 508
Loop β5-β6500 - 508
T R
elax
tion
time
in m
s2
120
100
20
Supplementary Figure 3. 15N T2 relaxation time measurement of the MED25 VBD. (a) 15N T2 relaxation times of the free MED25 VBD are plotted against the residue number. Error bars represent s.d. The N- and C-terminus and the long loop between 1 and 2 display significantly longer T2 times, indicating local flexibility. Due to line broadening and peak overlap, T2 times of the residues forming loop 5/ 6 could only be determined for T503. (b) The 25 lowest energy structures of the MED25 VBD are shown overlaid on the secondary structure elements. Loop
2/ 3 and loop 5/ 6 close the barrel from the bottom. Nature Structural & Molecular Biology doi:10.1038/nsmb.1999

: : : : | : : : : : : : : | : MED25 VBD GEFGQQSVSNKLLAWSGVLEWQEKPKPASVDANTKLTRSLPCQVYVNHGENLKTEQWPQKLIMQLIPQQLLTTLGPLFRNSRMVQFHFTNKDLESLKGLYRIMGNGFAGCVHFPHTAPCEVRVLMLLYSSKKKIFMGLIPYDQSGFVNGIRQVITNHKQVQQQKLKU70/PDB:1JEQ -YISK---TRKRALSRLKLKLN--------------DIVISVGIYNVQKA-----FDDPGLMLGFKPLVLL--KKHHY-LRPSLFVYPEEGSSTLFSALLIKCLEEVAALCRYT-RNIP-PYFVALVPQEEPPGFQLVFL-----PFAD----------------SPOC/PDB:1OW1 -PVDM-LLKKYPIVWQGLLALK--------------NDTAAVQLHFV--GNNVHRSLPLPLRIQRMRLEAQLEGARRMTDYCLLLALPCSQTE-SLKAFITYLQAQAAGIINVPNP-----YVLQIFPPCISPHLMIVIASV-----------------------KU80/PDB:1JEQ FVQR----RHSIH-WPCRLTIG-------------SNLSIRIAAYK--------SEGK-CFSVGFCKSSQVQ--RRFFMGNQVLKVFAAAAAV-ALSSLIHALDDDMVAIVRY--KRAN-PQVGVAFPHIKYECLVYVQL-----PFME----------------
: : : : | : : : : 100 : : : : | : MED25 VBD LLLLLLLLLLEEEEEEEEEEEELLLLLLLLLLLLLLEEEEEEEEEEELLLLLLHHHLLLEEEEEEEEHHHHHHHHHHHLLEEEEEEEELLLLHHHHHHHHHHHHHHEEEEEELLLLLLLLLLEEEEEELLLLLLEEEEEELLHHHHHHHHHHHHHHHHLLLLLLLKU70/PDB: 1JEQ -HHHH---LLLLLLEEEEEELL--------------LLEEEEEEELLLLL-----LLLLEEEEEEEEHHHL--LLLLL-LLLLEEEEELLLHHHHHHHHHHHHHHLEEEEEEEE-LLLL-LEEEEEEEELLLLEEEEEEL-----LLHH----------------SPOC/PDB: 1OW1 -LLLL-HHHHLLEEEEEEEEEL--------------LEEEEEEEEEE--ELHHHHHLLLLEEEEEEELLHHHHHHHHLLLEEEEEEEELHHHH-HHHHLHHHHHHLEEEEEEELLL-----EEEEEELLLLLLLEEEEEEEL-----------------------KU80/PDB: 1JEQ HHHH----LLLLL-EEEEEEEL-------------LLEEEEEEEEE--------LLLL-EEEEEEEEHHHLL--HHHLEEEEEEEEEELHHHH-HHHHHHHHHHHLEEEEEEE--LLLL-LEEEEEEEEELLEEEEEEEL-----LLHH----------------
α2α1 α3β1 β2 β3 β4 β5 β6 β7
10 20 9060504030 1301201101008070 140 150 160
Supplementary Figure 4. Structural homology search by DALI2. Alignment of amino acid sequences (upper panel) and structural elements (lower panel) with the three structural homologous proteins of MED25 VBD: KU70/PDB: 1JEQ3, SPOC/PDB: 1OW14, KU80/PDB:1JEQ3. Extended regions are highlighted in red and helical portions are shown in blue (lower panel).
Nature Structural & Molecular Biology doi:10.1038/nsmb.1999

a
2.00.0 3.01.0 4.0
Molar ratio
kcal
mol
e in
ject
ant
0.0
-0.5
K =1.6 μMd
-10
-8
-6-4
-2
0μ
cal s
ec
Time (min)0 20 40 60 80 100
K ~50 nMd
Molar ratio
kcal
mol
e in
ject
ant
-10-8-6-4-20
-12
μca
l sec
0.0
-0.2
-0.4
0 20 40 60 80 100Time (min)
2.00.0 3.01.0
b
-1-1
-1-1
Supplementary Figure 5. Isothermal calorimetry titration. (a) Isothermal calorimetry titration of MED25 VBD with VP16 TADn revealed a Kd of 1.6 μM. (b) Isothermal calorimetry titration of MED25 VBD with VP16 full-length TAD revealed a Kd of approximately 50 nM. The C-terminal portion of the VP16 TAD contributes to the binding to MED25 VBD and likely accounts for the difference in Kd between TADn and TAD.
Nature Structural & Molecular Biology doi:10.1038/nsmb.1999

9.5 9.0 8.5 8.0 7.5 7.0 6.5
130
125
120
115
110
105
9.5 9.0 8.5 8.0 7.5 7.0 6.5
130
125
120
115
110
105
++
WT VBD Q451E VBDN TAD15
N TAD15
N TAD15
15N
(pp
m)
15N
(pp
m)
15N
(pp
m)
H (ppm)1 H (ppm)1
9.5 9.0 8.5 8.0 7.5 7.0 6.5
130
125
120
115
110
105
H (ppm)1
a b c
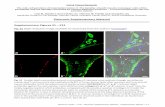
Supplementary Figure 6. The MED25 VBD Q451E mutation on 3 adjacent to the hydrophobic pocket also disrupts binding of full-length VP16 TAD to MED25 VBD. 1H-15N-HSQC spectra of free VP16 TAD (a), at 1:1 ratio of wild-type MED25 VBD (b) and with 1:1.5 excess of Q451E MED25 VBD (c) show that VP16 TAD only loosely binds, as seen by minor chemical shift changes, to mutant MED25 VBD without adopting a folded conformation. Far-shifted and broadened signals caused by the addition of wild-type MED25 VBD (b), are missing when the mutant MED25 VBD is added to 15N-labeled VP16 TAD (c).
Nature Structural & Molecular Biology doi:10.1038/nsmb.1999

9.5 9.0 8.5 8.0 7.5 7.0 6.5130
125
120
115
110
Phe442
Leu444
Gly448
Met447Asp441
Leu446
105
15N
(pp
m)
H (ppm)1
Asp445
Asp439
Supplementary Figure 7. 1H-15N-HSQC spectra of the VP16 TAD (black signals) and the VP16 TADn (red signals) both in the presence of 1.3 equivalents MED25 VBD. Far-shifted resonances overlap for both peptides and are indicated by circles and assignments.
Nature Structural & Molecular Biology doi:10.1038/nsmb.1999

10 9 8 7
130
125
120
115
110
Gly524
Leu525
Gln456
Ile521
Val471
TADc
TADc
TADn
1 H (ppm)
15N
(pp
m)
Gln456
Gln456Ile521
Ile521
V471
Leu525
Gly524
a
b
β4
β7α1
β6
β4 β7
α3
β6
α3α2
α1
α2
β3
β5
β5β1
105
Supplementary Figure 8. Mapping the TADc binding site on MED25 VBD. (a) Overlay of 1H-15N-TROSY-HSQC5 spectra of the MED25 VBD bound to the VP16 TADn (red) and the MED25 VBD bound to the VP16 TAD (black). (b) A cartoon representation of MED25 VBD shows the clustering of the residues experiencing distinct chemical shift changes upon interaction with VP16 TADc on 4 and 7. These residues are highlighted in blue. The TADn and TADc binding sites are located on different sides of the MED25 VBD barrel.
Nature Structural & Molecular Biology doi:10.1038/nsmb.1999

WT K447E Q451E H499E K545E
anti-Flag
Supplementary Figure 9. Wild-type and mutant MED25 VBD displayed comparable level of expression in transfected HEK293T cells when immunoblotted with anti-Flag antibody.
Nature Structural & Molecular Biology doi:10.1038/nsmb.1999

Supplementary Table 1. Ramachandran Plot Summary of the MED25 VBD structure from PROCHECK6.
Ramachandran Plot Summary from PROCHECK
Most favoured regions 89.1%
Additionally allowed regions 10.4%
Generously allowed regions 0.5%
Disallowed regions 0.0%
Supplementary References 1. Gouet, P., Courcelle, E., Stuart, D.I. & Metoz, F. ESPript: analysis of multiple sequence
alignments in PostScript. Bioinformatics 15, 305-308 (1999). 2. Holm, L. & Sander, C. Protein structure comparison by alignment of distance matrices. J
Mol Biol 233, 123-138 (1993). 3. Walker, J.R., Corpina, R.A. & Goldberg, J. Structure of the Ku heterodimer bound to
DNA and its implications for double-strand break repair. Nature 412, 607-614 (2001). 4. Ariyoshi, M. & Schwabe, J.W. A conserved structural motif reveals the essential
transcriptional repression function of Spen proteins and their role in developmental signaling. Genes Dev 17, 1909-1920 (2003).
5. Pervushin, K., Riek, R., Wider, G. & Wüthrich, K. Attenuated T2 relaxation by mutual cancellation of dipole-dipole coupling and chemical shift anisotropy indicates an avenue to NMR structures of very large biological macromolecules in solution. Proc Natl Acad Sci U S A 94, 12366-12371 (1997).
6. Laskowski, R.A., Rullmannn, J.A., MacArthur, M.W., Kaptein, R. & Thornton, J.M. AQUA and PROCHECK-NMR: programs for checking the quality of protein structures solved by NMR. J Biomol NMR 8, 477-486 (1996).
Nature Structural & Molecular Biology doi:10.1038/nsmb.1999