Scientific Workflows - University of...
Transcript of Scientific Workflows - University of...

Scientific Workflows, B. LudäscherScientific Workflows, B. LudäscherpPOD @ NESCENT, Sept ’07pPOD @ NESCENT, Sept ’07
Scientific Workflows: A(nother) Vision of
pPOD “Data Integration” !?
Bertram LudäscherShawn BowersTimothy McPhillipsDave Thau
UC DAVISDepartment ofComputer Science
Dept. of Computer Science & UC Davis Genome Center
University of California, DAVIS

Scientific Workflows, B. LudäscherScientific Workflows, B. LudäscherpPOD @ NESCENT, Sept ’07pPOD @ NESCENT, Sept ’07
Overview
• Scientific Workflow: – Overview Vision– Examples using Kepler (from NSF/ITR SEEK)
• Provenance in Scientific Workflows– from single runs to project histories
• pPOD & Kepler– next steps

Scientific Workflows, B. LudäscherScientific Workflows, B. LudäscherpPOD @ NESCENT, Sept ’07pPOD @ NESCENT, Sept ’07
Different Kinds of (Data) “Integration” • “Traditional” Information (& Data) Integration
– syntactic & structural heterogeneities, schema mappings, schema matching, query rewriting (parsing, matching, [G]LAV, Chase [+IC], Resolution), …
– dealing with fundamentally same (largely overlapping) information– find ways to integrate different representations
• Scientific Information Integration (SII)– includes the above– … but often deals with combining fundamentally different information – more than one way to combine, “integrate” the data – integration invokes scientific theories, models that cannot be
inferred from only data, schema, ontologies
“joining” of data, “chaining” of analysis steps in the scientist’s head ( … y := f(x) ; z := g(x,y); … ) – make these analysis pipelines first-class citizens– scientific workflows can provide an end-to-end framework

Scientific Workflows, B. LudäscherScientific Workflows, B. LudäscherpPOD @ NESCENT, Sept ’07pPOD @ NESCENT, Sept ’07
Data Source
Data Source
Data Source
Local schema Local schema Local schema
Component schema Component schema Component schema
Export schema Export schema Export schema
Federated schema Federated schema
Export schema Export schema
Types of “Information Integration”• Conventional information integration:
– schema-based – view-based – at the instance level
• Spatial (co-)registration/“overlay” of different data– from 2D, 3D, 4D (x,y,z,t), (4+n) D GIS ++
• Extended DI approaches using “ontologies”– controlled vocabularies, metadata, annotations
• Scientific Information Integration= data + process/application integration scientific workflows
• … can include all the others and – …statistics, data mining, visualization, …

Scientific Workflows, B. LudäscherScientific Workflows, B. LudäscherpPOD @ NESCENT, Sept ’07pPOD @ NESCENT, Sept ’07
Scientific Workflows = Cyberinfrastructure UPPER-WARE
Science Environment for Ecological
Knowledge (“SEEK”)
Underware
Middleware
UpperMiddleware
Upperware

Scientific Workflows, B. LudäscherScientific Workflows, B. LudäscherpPOD @ NESCENT, Sept ’07pPOD @ NESCENT, Sept ’07
Science Environment for Ecological Knowledge (SEEK)Access distributed environmental, ecological, and systematics data
– Enable data sharing & reuse– Enhance data discovery at global scales– Distributed data network
EcoGrid
Design, reuse, and execute scientific analyses – Enable communication and collaboration for analysis– Enable reuse of analytical components and analyses– Integrated data access
Kepler
Data discovery and integration– Addressing variety of semantic data heterogeneity issues– Ontology and controlled-vocabulary development– Semantic data and actor annotations– Resolve taxonomic ambiguities
SMS / OBOE / Taxonomic concept services

Scientific Workflows, B. LudäscherScientific Workflows, B. LudäscherpPOD @ NESCENT, Sept ’07pPOD @ NESCENT, Sept ’07
Kepler Data Access via the EcoGrid
• Lightweight API for providers & clients• Implemented via web services • Common metadata query syntax• Common mechanism for accessing ecological (KNB), museum specimen (DiGIR), environmental (SRB), and geological (GEON) data
• “Catalog-based Integration”• NOT a single CDM• leave the integration to the workflow designer!

Scientific Workflows, B. LudäscherScientific Workflows, B. LudäscherpPOD @ NESCENT, Sept ’07pPOD @ NESCENT, Sept ’07
Scientific Workflow
Capture how a scientist works with data and analytical tools– data access, transformation, analysis, visualization– possible worldview: dataflow-oriented
Scientific workflow (wf) benefits (compare w/ script-based approaches) : – wf automation – wf & component reuse – wf design, documentation– wf archival, sharing– built-in concurrency
(task-, pipeline-parallelism) – built-in provenance support– distributed execution
(Grid) support – …

Scientific Workflows, B. LudäscherScientific Workflows, B. LudäscherpPOD @ NESCENT, Sept ’07pPOD @ NESCENT, Sept ’07
Kepler Collaboration (alive and evolving)• Open-source
– Builds on Ptolemy II from UC Berkeley
• Contributors from:– SEEK– SciDAC SDM– Ptolemy– GEON– ROADNet– Resurgence– AToL: CIPRES, POD– …
• Goals– Create powerful analytical
tools that are useful across disciplines
– Ecology, Biology, Engineering, Geology, Physics, Chemistry, Astronomy, …
Ptolemy IIPtolemy II
Phyl-O'Data (POD)
Natural Diversity
Discovery Project

Scientific Workflows, B. LudäscherScientific Workflows, B. LudäscherpPOD @ NESCENT, Sept ’07pPOD @ NESCENT, Sept ’07
Basic Kepler User Interface
WorkflowCanvas
Actor Libraries
ThumbnailNavigation
QuickSearch
Tool Bar

Scientific Workflows, B. LudäscherScientific Workflows, B. LudäscherpPOD @ NESCENT, Sept ’07pPOD @ NESCENT, Sept ’07
Kepler Data Access via the EcoGrid
Data QuickSearch Tab
Metadata Keyword Search
Access Multiple EcoGrid Sources
Return Data Setsas “Actors” to
Drag-Drop to Canvas

Scientific Workflows, B. LudäscherScientific Workflows, B. LudäscherpPOD @ NESCENT, Sept ’07pPOD @ NESCENT, Sept ’07
Input/Output Semantic Annotation
Actor input/output port annotation:– Each port can be annotated
with multiple classes from multiple ontologies
– Annotations are stored with actor metadata (MOML)
– Actors can be discovered, validated, etc., via their “semantic types”

Scientific Workflows, B. LudäscherScientific Workflows, B. LudäscherpPOD @ NESCENT, Sept ’07pPOD @ NESCENT, Sept ’07
Actor Annotations
• Actor Annotations for Indexing & Classification
– New actors can be annotated and indexed into the component library (e.g., specializing generic actors)
– Existing components can also be revised, annotated, and indexed (hiding previous versions)
– Quick search leverages metadata, including annotations & ontologies

Scientific Workflows, B. LudäscherScientific Workflows, B. LudäscherpPOD @ NESCENT, Sept ’07pPOD @ NESCENT, Sept ’07
Kepler Demo: Building a simple workflow
Select actors from Kepler actor library:– Local or remote actors– View actor metadata/documentation (not shown)– Drag desired actor to canvas– Connect actor ports
other actor examples
1
2
3

Scientific Workflows, B. LudäscherScientific Workflows, B. LudäscherpPOD @ NESCENT, Sept ’07pPOD @ NESCENT, Sept ’07
Kepler Demo: Building a simple workflow
Select input data:– Shown here is an EcoGrid for “bacterial abundance”– Connect data “actors” to workflow inputs
many ways to import data
3
1
2

Scientific Workflows, B. LudäscherScientific Workflows, B. LudäscherpPOD @ NESCENT, Sept ’07pPOD @ NESCENT, Sept ’07
Kepler Demo: Building a simple workflow
Using EcoGrid data sources:– Display metadata (EML)– Query data via SQL/QBE interface– … even if it is a tab-delimited file (see
above)

Scientific Workflows, B. LudäscherScientific Workflows, B. LudäscherpPOD @ NESCENT, Sept ’07pPOD @ NESCENT, Sept ’07
Kepler Demo: Building a simple workflow
Run the workflow …– Also set parameters, select &
configure director, run window, etc.

Scientific Workflows, B. LudäscherScientific Workflows, B. LudäscherpPOD @ NESCENT, Sept ’07pPOD @ NESCENT, Sept ’07
SEEK Ecological Niche Modeling WorkflowsComplex workflows with many levels of nesting (sub-workflows)
– Predict species locations from presence data and environmental layers– Designed to support different prediction algorithms (reusability)– Currently uses GARP (Genetic Algorithm for Rule-Set Prediction)
n levels down

Scientific Workflows, B. LudäscherScientific Workflows, B. LudäscherpPOD @ NESCENT, Sept ’07pPOD @ NESCENT, Sept ’07
Drilling down: Calculate Best Rulesets
climate change data

Scientific Workflows, B. LudäscherScientific Workflows, B. LudäscherpPOD @ NESCENT, Sept ’07pPOD @ NESCENT, Sept ’07
SEEK Ecological Niche Modeling Workflows• Includes a number of workflows for automating “special purpose”
data-integration tasks– Integration of multiple data sets and data types– Workflows for local caching of data, format and content conversions
Rescale grid data, adjust resolutions, extents, merges grids
Integrate Hydro1K North and South American data, including warp/projection, format conversion, rescaling, etc.

Scientific Workflows, B. LudäscherScientific Workflows, B. LudäscherpPOD @ NESCENT, Sept ’07pPOD @ NESCENT, Sept ’07
The Joy of Exa-Scale Cyberinfrastructure
• Are we working at the right level of abstraction?Are we working at the right level of abstraction?• Are we optimizing the right thing?Are we optimizing the right thing?• Optimize human cycles, not just CPU cycles!Optimize human cycles, not just CPU cycles!
– cf. John McCarthy (of AI/LISP fame) cf. John McCarthy (of AI/LISP fame)
Make data & scientific workflows effectively (re-)usable Make data & scientific workflows effectively (re-)usable for scientistfor scientist
Make workflows first-class, shareable “knowledge Make workflows first-class, shareable “knowledge artifacts”artifacts”
Support user-oriented provenance queriesSupport user-oriented provenance queries

Scientific Workflows, B. LudäscherScientific Workflows, B. LudäscherpPOD @ NESCENT, Sept ’07pPOD @ NESCENT, Sept ’07
(Data) Provenance & Scientific Workflows• (Data) provenance
– data lineage, processing history
• Query the lineage of a data product: – what data it is derived from and how
• Evaluate the results of a workflow: – is the approach correct
• Reuse intermediate or final products of one workflow in another
• Explain unexpected results• Discover all results derived from a given data set• Accurately prepare methods section of a
publication• Archive scientific results in a repository• Replicate the results reported by another researcher

Scientific Workflows, B. LudäscherScientific Workflows, B. LudäscherpPOD @ NESCENT, Sept ’07pPOD @ NESCENT, Sept ’07
Inferring a phylogenetic tree from disparate data
Actors
Maximum likelihood tree
(DNA)
Maximum parsimony tree
Maximum likelihood tree (continuous characters)
Aligned DNA sequences
Discrete morphological
data
Continuous characters
Consensus Tree(s)
“Integrate”
Datasets Datasets
ProvenanceStore

Scientific Workflows, B. LudäscherScientific Workflows, B. LudäscherpPOD @ NESCENT, Sept ’07pPOD @ NESCENT, Sept ’07
“Scientific” provenance questions (single run)• What DNA sequences were input (phylogenetic trees
were output) by the workflow?
• What intermediate phylogenetic trees were created?
• Which actor created this phylogenetic tree?
• Which input sequences does this consensus tree depend on?
• Which input sequences were not used to derive any consensus tree
• What sequence alignment (key intermediate data) was used to infer this tree?

Scientific Workflows, B. LudäscherScientific Workflows, B. LudäscherpPOD @ NESCENT, Sept ’07pPOD @ NESCENT, Sept ’07
A (very) simple phylogenetics workflow

Scientific Workflows, B. LudäscherScientific Workflows, B. LudäscherpPOD @ NESCENT, Sept ’07pPOD @ NESCENT, Sept ’07
TextFileReader:1NexusFileParser:1
PhylipConsense:1
PhylipPars:
1
PhylipPars:3
PhylipPars:5
NexusFileParser:1
PhylipPars:1
PhylipPars:1
PhylipPars:1PhylipPars:1
PhylipConsense:1
PhylipConsense:1
PhylipConsense:1
PhylipConsense:1
PhylipConse
nse:1
Phyl
ipCo
nsen
se:1
• Derivation (processing history) of a data item in a scientific workflow run (a DAG)– Nodes = data items the workflow run operated on or created– Edges = “was directly used in”
• … labeled by the actor invocation that performed this computation
• Different (emerging) provenance extensions to Kepler
Data lineage + processing history for a consensus tree

Scientific Workflows, B. LudäscherScientific Workflows, B. LudäscherpPOD @ NESCENT, Sept ’07pPOD @ NESCENT, Sept ’07
Provenance: Single Run

Scientific Workflows, B. LudäscherScientific Workflows, B. LudäscherpPOD @ NESCENT, Sept ’07pPOD @ NESCENT, Sept ’07
Provenance: Multiple Runs

Scientific Workflows, B. LudäscherScientific Workflows, B. LudäscherpPOD @ NESCENT, Sept ’07pPOD @ NESCENT, Sept ’07
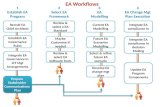
Conceptual workflows: series of subworkflows

Scientific Workflows, B. LudäscherScientific Workflows, B. LudäscherpPOD @ NESCENT, Sept ’07pPOD @ NESCENT, Sept ’07
Manual, data visualization, and quality assessment steps are interleaved with automated steps

Scientific Workflows, B. LudäscherScientific Workflows, B. LudäscherpPOD @ NESCENT, Sept ’07pPOD @ NESCENT, Sept ’07
Projects comprise multiple
conceptual workflows

Scientific Workflows, B. LudäscherScientific Workflows, B. LudäscherpPOD @ NESCENT, Sept ’07pPOD @ NESCENT, Sept ’07
Workflows are run multiple times with different parameter settings

Scientific Workflows, B. LudäscherScientific Workflows, B. LudäscherpPOD @ NESCENT, Sept ’07pPOD @ NESCENT, Sept ’07
‘Aware’ of only one workflow, one run at a time
Data, workflows, and provenance records reside outside the system between runs
Users must perform most data and provenance management outside of the system
Workflows must be modified or reconfigured to operate on different input data
How Kepler is used today
• p1• p2• p3
• p1• p2• p3

Scientific Workflows, B. LudäscherScientific Workflows, B. LudäscherpPOD @ NESCENT, Sept ’07pPOD @ NESCENT, Sept ’07
•Data is registered •Project folders allow users to organize data.•Project history records and depicts past workflow runs and the flow of data between runs.•Data is staged from the project folders (and project history).•Run outputs appear in the project history (along with the input) if the run is committed.•All or part of the output of a run may be used to update the project folders.•Workflows can be applied to different data sets without modifying their definitions.
Support for project folders & histories

Scientific Workflows, B. LudäscherScientific Workflows, B. LudäscherpPOD @ NESCENT, Sept ’07pPOD @ NESCENT, Sept ’07
Recomputed data can replace old versions, be stored elsewhere in folders, or simply left in the project history.
Replaced data are always accessible via project history.
Provenance queries provide access to all data regardless of location.
Project history relieves need to perform data versioning via project
folders

Scientific Workflows, B. LudäscherScientific Workflows, B. LudäscherpPOD @ NESCENT, Sept ’07pPOD @ NESCENT, Sept ’07
Workflow library is not a flat list of available workflows.
Workflows evolve throughout a project, and previous versions must be retained for reference and for further use.
Workflow evolution view complements run history.
Managing workflow evolution

Scientific Workflows, B. LudäscherScientific Workflows, B. LudäscherpPOD @ NESCENT, Sept ’07pPOD @ NESCENT, Sept ’07
Summary & Next Steps• Kepler today
– used in ecoinformatics (SEEK), ChIP-chip, geoinformatics, …– data catalog, data grid– workflows for data integration– data annotation and semantic extensions
• Kepler next steps (planned deliverables):– PHYLOGENETIC SCIENTIFIC WORKFLOWS
– Develop use cases / conceptual workflows:– tree construction (understood)– post-tree analysis, supertree/matrix construction (exciting :)
community-driven!– Implement subset of those in Kepler– Generate actor library targeting community use cases
– PROJECT HISTORIES SUPPORT (cf. DILS'07 paper)– Extend use cases to exploit project histories / provenance– Implement those
– pPOD “REPOSITORY” (Orchestra!?)1. Extend Kepler to use pPOD data repository

Scientific Workflows, B. LudäscherScientific Workflows, B. LudäscherpPOD @ NESCENT, Sept ’07pPOD @ NESCENT, Sept ’07
Consilience: The Unity of Knowledge (E. O. Wilson)• "Literally a jumping together of
knowledge by the linking of facts and fact-based theory across disciplines to create a common groundwork for explanation."
– E.O.Wilson
• eScience, Cyberinfrastructure: mechanisms to make progress
• Scientific Workflows: crucial elements to get the most mileage out of CI to fuel eScience, accelerating knowledge discovery
• Identify the real bottlenecks in this quest!
Wer Visionen hat, sollte zum Arzt gehen – Helmut Schmidt on Willy Brandt
We must know, we will know.-- David Hilbert

Scientific Workflows, B. LudäscherScientific Workflows, B. LudäscherpPOD @ NESCENT, Sept ’07pPOD @ NESCENT, Sept ’07
kepler-project.org
Questions …

Scientific Workflows, B. LudäscherScientific Workflows, B. LudäscherpPOD @ NESCENT, Sept ’07pPOD @ NESCENT, Sept ’07
References
• Niche Modeling– D Pennington, D Higgins, AT Peterson, M Jones, B
Ludaescher, S Bowers. Ecological niche modeling using the Kepler workflow system.. Workflows for e-Science: Scientific Workflows for Grids, Springer-Verlag, 2007.
– Ecological Niche Modeling in Kepler. User Manual. Draft, 2007
• Semantic Annotation– S Bowers, B Ludaescher. A calculus for propagating semantic
annotations through scientific workflow queries. QLQP, 2006.– S Bowers, B Ludaescher. Actor-oriented design of scientific
workflows. ER, 2005.– C Berkley, S Bowers, M Jones, B Ludaescher, M Schildhauer, J
Tao. Incorporating semantics in scientific workflow authoring. SSDBM, 2005.
– S Bowers, B Ludaescher. An Ontology-driven framework for data transformation in scientific workflows. DILS, 2004.

Scientific Workflows, B. LudäscherScientific Workflows, B. LudäscherpPOD @ NESCENT, Sept ’07pPOD @ NESCENT, Sept ’07
References
• Provenance in Workflows– S Bowers, T McPhillips, M Wu, B Ludaescher. Project
histories: Managing data provenance across collection-oriented scientific workflow runs. DILS, 2007.
– S Bowers, T McPhillips, B Ludaescher. Provenance in collection-oriented workflows. Concurrency and Computation: Practice and Experience, 2007.
– B Ludaescher, N Podhorszki, I Altintas, S Bowers, T McPhillips. From computation models to models of provenance: The RWS approach. Concurrency and Computation: Practice and Experience, 2007.

Scientific Workflows, B. LudäscherScientific Workflows, B. LudäscherpPOD @ NESCENT, Sept ’07pPOD @ NESCENT, Sept ’07
Additional Related PublicationsSemantic Type Annotation
– S Bowers, B Ludaescher. A Calculus for Propagating Semantic Annotations through Scientific Workflow Queries. ICDE Workshop on Query Languages and Query Processing (QLQP), LNCS, 2006.
– S Bowers, B Ludaescher. Towards Automatic Generation of Semantic Types in Scientific Workflows. International Workshop on Scalable Semantic Web Knowledge Base Systems (SSWS), WISE 2005 Workshop Proceedings, LNCS, 2005.
– C Berkley, S Bowers, M Jones, B Ludaescher, M Schildhauer, J Tao. Incorporating Semantics in Scientific Workflow Authoring. SSDBM, 2005.
– B Ludaescher, K Lin, S Bowers, E Jaeger-Frank, B Brodaric, C Baru. Managing Scientific Data: From Data Integration to Scientific Workflows. GSA Today, Special Issue on Geoinformatics, 2006.
– S Bowers, D Thau, R Williams, B Ludaescher. Data Procurement for Enabling Scientific Workflows: On Exploring Inter-Ant Parasitism. VLDB Workshop on Semantic Web and Databases (SWDB), 2004.
– S Bowers, K Lin, B Ludaescher. On Integrating Scientific Resources through Semantic Registration. SSDBM, 2004. – S Bowers, B Ludaescher. An Ontology-Drive Framework for Data Transformation in Scientific Workflows. International Workshop on
Data Integration in the Life Sciences (DILS), LNCS, 2004. – S Bowers, B Ludaescher. Towards a Generic Framework for Semantic Registration of Scientific Data. International
Semantic Web Conference Workshop on Semantic Web Technologies for Searching and Retrieving Scientific Data, 2003.
Workflow Design and Modeling– T McPhillips, S Bowers, B Ludaescher. Collection-Oriented Scientific Workflows for Integrating and Analyzing Biological
Data. Workshop on Data Integration in the Life Sciences (DILS), LNCS, 2006.– S Bowers, T McPhillips, B Ludaescher, S Cohen, SB Davidson. A Model for User-Oriented Data Provenance in Pipelined
Scientific Workflows. International Provenance and Annotation Workshop (IPAW), LNCS, 2006.– S Bowers, B Ludaescher, AHH Ngu, T Critchlow. Enabling Scientific Workflow Reuse through Structured Composition of
Dataflow and Control-Flow. IEEE Workshop on Workflow and Data Flow for Scientific Applications (SciFlow), 2006.– S Bowers, B Ludaescher. Actor-Oriented Design of Scientific Workflows. International Conference on Conceptual Modeling
(ER), LNCS, 2005.– T McPhillips, S Bowers. Pipelining Nested Data Collections in Scientific Workflows. SIGMOD Record, 2005.
Kepler – D Pennington, D Higgins, AT Peterson, M Jones, B Ludaescher, S Bowers. Ecological Niche Modeling using the Kepler
Workflow System. Workflows for e-Science, Springer-Verlag, to appear.– W Michener, J Beach, S Bowers, L Downey, M Jones, B Ludaescher, D Pennington, A Rajasekar, S Romanello, M Schildhauer, D
Vieglais, J Zhang. SEEK: Data Integration and Workflow Solutions for Ecology. Workshop on Data Integration in the Life Sciences (DILS), LNCS, 2005.
– S Romanello, W Michener, J Beach, M Jones, B Ludaescher, A Rajasekar, M Schildhauer, S Bowers, D Pennington. Creating and Providing Data Management Services for the Biological and Ecological Sciences: Science Environment for Ecological Knowledge. SSDBM, 2005.