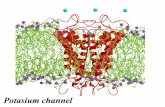
Potassium Channels
description
Transcript of Potassium Channels

Doyle et al. (1998) Science 280:69-77
Missy CavallinSeptember 14, 2007
Potassium Channels

Outline
Types of K+ channels Voltage-gated
Functional roles Nomenclature Structure Activation Inactivation
Assigned Experimental Papers (structure, voltage sensor, inactivation)

Armstrong and Hille (1998) Neuron 20:371-380
Types of K+ Channels
• Inward Rectifying• Ca+2 sensitive• ATP-sensitive• Na+ activated• Cell volume sensitive• Type A• Receptor-coupled• Voltage-gated

Inward Rectifying (KIR
)
2 transmembrane regions (M1 and M2) 4 subunits form pore P region between M1 and M2 Regulated by concentration of extracellular
potassium Activity dependent on interactions with
phosphatidylinositol 4,5-bisphosphate (PIP2) Blocked by external Ba+

Functions of KIR
Maintain membrane resting potential near EK
non-conducting at positive membrane potentials Contributes to cell excitability

Ca+2 sensitive
4 protein subunits Selectivity filter on external surface RCK domains act as gate
2 Ca+2 ions bind to RCK to regulate gate 3 types of channels
High conductance (BK): 100-220 pS Intermediate conductance (IK): 20-85 pS Small conductance (SK): 2-20 pS

Functions of Ca+2 sensitive
Generate membrane potential oscillations Afterhyperpolarization

ATP-sensitive
2 Transmembrane regions 4 subunits Inhibited by ATP Inwardly rectifying Not voltage dependent

Functions of ATP-sensitive
Couples K+ conductance to cell metabolic state Responds to metabolic changes: e.g. senses
glucose concentration in -cells Senses intracellular nucleotide concentrations

Na+ activated
Voltage-insensitive Blocked by Mg+2 and Ba+2

Cell volume sensitive
Activated by increase in cell volume Blocked by quinidine, lidocaine, cetiedil

Type A
Tetramer of -subunits and intracellular -subunits
Rapid activation and inactivation Inactivation may involve -subunits
Possible role in delaying spikes by regulation of fast phase of action potentials

Receptor-coupled
Muscarinic-inactivated Slow activation Non-inactivating Non-rectifying
Atrial muscarinic-activated Inward rectifying

Voltage-gated channels: Function
Regulate resting membrane potential toward EK
Control shape and frequency of action potentials Keep fast action potentials short Terminate intense periods of activity Time interspike intervals
Lower the effectiveness of excitatory inputs on a cell when open
Delayed rectifier type expressed in axons Delayed activation and slow inactivation (shape
action potential)

Gutman et al. (2005) Pharmacol Rev 57:473-508
Nomenclature of voltage-gated K+ channels

Gulbis et al (2000) Science 289:123-127
Voltage-gated: Structure
6 transmembrane (TM) regions (S1-S6)
Principal subunits Homo- or
heterotetramers S5 & S6 of each
subunit surround pore Auxiliary subunits
Regulate channel activity

More Structure
S4 segment has multiple positively charged amino acids (arginine or lysine) voltage sensor
P loop between S5 & S6 (selectivity filter) G-Y-G sequence required for K+ selectivity
T1 domain: tetramerization domain; connects subunits to subunits

Doyle et al. (1998) Science 280:69-77
K+ movement through channel
K+ attracted to negative charge of helices near selectivity filter
K+ becomes hydrated before exiting channel

Voltage-gated: Activation
Activated by depolarization Voltage sensor (S4) Translocation of charges on S4
Helical screw Lateral movement of crossed helices Rocking motion at interface of 2 domains

Kurata and Fedida (2006) Progress in Biophysics and Molecular Biology 92:185-208
Voltage-gated: Inactivation
N-type: ball-and-chain Amino acids at N-terminus occlude intracellular
channel pore Rapid inactivation
C-type Conformational changes at selectivity filter or
extracellular entrance to channel Slow inactivation

Rasmusson et al (1998) Circ. Res. 82:739-750

Paper 1
Crystal Structure of a Mammalian Voltage-Dependent Shaker Family K+ Channel
Stephen B. Long, Ernest B. Campbell, and Roderick MacKinnon
Science (2005) 309:897-903

Methods
Kv1.2 with 2 subunit from rat brain expressed
in yeast X-ray crystallography 2.9 Å resolution

Figure 1 A) Electron density map = blue mesh; final model trace = yellow B) Crystal lattice structure of channel: transmembrane segments of subunit = red; subunit + T1 = blue; unit cell = black box

Kv1.2 structural features
4-fold symmetry (tetrad axis of unit cell) Dimensions of tetramer: ~135 Å x 95 Å x 95 Å Length of transmembrane segments: ~ 30 Å
(approximately thickness of lipid bilayer)

Figure 2 A) Side view of ribbon model of channel with 4 subunits of unique color. NADP+ cofactor = black sticks B) 1 subunit from panel A illustrating S1-S6 as well as N- and C-termini C) Looking into pore from extracellular side to show interactions of 4 subunits.

Figure 3 A) Side view of 2 subunits of three voltage-gated K channels: Kv1.2 = red; KcsA = gray; KvAP = blue; outer = S5 of Kv1.2; inner = S6 of Kv1.2 B) Top view into poreNote the overlap of the structures (conserved).

Paper 2
How Does a Voltage Sensor Interact with a Lipid Bilayer? Simulations of a Potassium Channel
Domain
Zara A. Sands and Mark S.P. Sansom
Structure (2007) 15:235-244

How is a positively charged S4 helix able to stably span the lipid bilayer?
Used molecular dynamics simulations to test the interactions of the voltage sensor domain of KvAP (archaebacterial) with an artificial phosphatidylcholine (PC) bilayer or with a more natural PC + phosphatidylglycerol (PG) bilayer Computer modeling Extended time (50 ns) vs. other models (20 ns) to
increase sampling

Results
Penetration of water into center of voltage sensor and bilayer
Local deformation of bilayer due to interactions of lipid head groups with arginine side chains of S4 subunit Electrostatic field is focused at center of bilayer
Numerous hydrogen bonds between phosphate head groups of lipids with arginine residues of S4 subunit

Figure 1 Lipids interact strongly with and are drawn into voltage sensor causing changes in PC bilayer conformation. A) Voltage sensor shown as ribbon (S1-3 = gray; S4 = magenta); lipid phosphate head groups colored based on z coordinate (red = extracellular; blue = intracellular). Arrow shows phosphate that is pulled away from surface. B) Arginines on S4 interactions with lipid phosphate groups.

Figure 2 Shows lipid bilayer compression when voltage sensor present vs. respected control bilayers without proteins. (distance between upper and lower phosphate atoms)

Figure 3 There are more hydrogen bonds formed in PC bilayer. Although both bilayers have hydrogen bonds between S4 arginines and lipid phosphate groups or water.

Figure 4 A and C) red = water; blue = lipid B) blue = cationic side chain; red = anionic side chain
PC/PG overall has more hydrogen bonds.
Increased hydrogen bonds at termini may stabilize charged S4 in bilayer.
Lack of water to hydrogen bonds at R133 indicates prevention of water penetration.

Figure 6 A) Water in cavities of of voltage sensor but not at constriction point. B) Model of average pore radius. C) Electrostatic potential distribution (± 120 mV). Note the focus of electrical potential around salt bridge at constriction point.

Conclusions
Hydrogen bonding may help to stabilize voltage sensor in membrane
Compression of bilayer can decrease the distance necessary for the movement of C-type inactivation of potassium channels

Paper 3
Slow Inactivation in Voltage Gated Potassium Channels is Insensitive to the Binding of Pore
Occluding Peptide Toxins
Carolina Oliva, Vivian Gonzalez, and David Naranjo
Biophysics Journal (2005) 89: 1009-1019

Toxin binding to K channels and Slow (C-type) Inactivation
Sensitive to K+ or tetraethylammonium (TEA) in pore
Mutations of external vestibule alter both processes
Does toxin binding interfere with C-type inactivation?

Figure 1 B) Amino acids important for toxin binding (left) and slow inactivation (right) overlap. red = scorpion toxin; yellow = conotoxin; orange = both; blue = slow inactivation

Methods
Oocytes Whole cell recording Outside-out patch recording
Toxins Conotoxin (-PVIIA) Charybdotoxin (CTX)

Results
Rate of inactivation and recovery are toxin insensitive
Inactivation does not alter toxin binding site

Figure 2 Whole cell voltage clamp. A) Conotoxin (thick trace) decrease current amplitude and causes slight delay in activation. B) Time course of ratio of currents in panel A. Toxin binding (and current ratio) reached steady state within recording interval.

Figure 3 and Table 1 show that the inactivation kinetics are not affected by conotoxin in spite of change in current amplitude. Therefore, conotoxin binding is independent of slow inactivation. Similar results were shown for CTX (Fig. 6). There are differences between whole cell vs. outside out patches, but not with regards to toxin effects.

Figure 5 A) Conotoxin decreases current amplitude in response to voltage steps. B) IV and GV relationships. Voltage shift does not change in the presence of toxin (bottom). V = -18 mV for no toxin; V = -21 mV with toxin

Conclusions
Inactivation kinetics were not affected by toxin binding
Bound toxin does not hinder conformation change involved with slow inactivation