Optimization of Protein Force-Field Parameters€¦ · Optimization of Protein Force-Field...
Transcript of Optimization of Protein Force-Field Parameters€¦ · Optimization of Protein Force-Field...
-
Optimization of Protein Force-Field Parameters
Yuko OKAMOTO (岡本 祐幸) Department of Physics and
Structural Biology Research Center
Graduate School of Science
and Center for Computational Science
Graduate School of Engineering
and Information Technology Center
NAGOYA UNIVERSITY (名古屋大学) e-mail: okamoto{a}phys.nagoya-u.ac.jp
URL: http://www.tb.phys.nagoya-u.ac.jp/
Seminar at the Basque Center for Applied Mathematics
July 14, 2014
-
cano
E
PB(E) = n(E)WB(E)
Canonical Probability Distribution
E
WB(E) = exp(- E )
Boltzmann Factor
E
n(E)
Density of States
Canonical Ensemble at
Temperature T
-
SA-2
P B
(E)
E E
P B
(E) = n(E)W B
(E)
High T
P B
(E)
E
E min
Low T
Canonical Distributions of Potential Energy
Intermediate T
-
30
20
10
0
-10
E
200000150000100000500000
MC Sweeps
Canonical 1000K
-
30
20
10
0
-10
E
200000150000100000500000
MC Sweeps
Canonical 600K
-
30
20
10
0
-10
E
200000150000100000500000
MC Sweeps
Canonical 50K
-
Generalized-Ensemble Algorithm(拡張アンサンブル法) Generic Term for Simulation Methods that Greatly Enhance
Conformational Sampling [e.g., Multicanonical Algorithm, Wang-Landau, Simulated Tempering, Replica-Exchange Method, etc.]
Based on Non-Boltzmann Weight Factors
Realize random walks in potential energy and/or any other physical quantities (OR their conjugate parameters)
Histogram Reweighting Techniques
Can obtain thermodynamic quantities for a wide range of temperature
and/or other parameter values from a single simulation run REVIEWS: U.H.E. Hansmann & Y.O., in Ann. Rev. Comput. Phys. VI, D. Stauffer (ed.) (World Scientific, Singapore, 1999) pp. 129-157;
A. Mitsutake, Y. Sugita, & Y.O., Biopolymers 60, 96 (2001); Y.O., J. Mol. Graphics Modell. 22, 425 (2004);
Y. Sugita, A. Mitsutake, & Y.O., in Lecture Notes in Physics,
W. Janke (ed.) (Springer-Verlag, Berlin, 2008) pp. 369-407; H. Okumura, S.G. Itoh, & Y.O., in Practical Aspects of Computational Chemistry II: An Overview of the Last Two Decades and Current Trends,
J. Leszczynski and M.K. Shukla (eds.) (Springer, Dordrecht, 2012) pp. 69-101;
A. Mitsutake, Y. Mori, and Y.O, in Biomolecular Simulations: Methods and Protocols,
L. Monticelli and E. Salonen (eds.) (Humana Press, New York, 2012) pp. 153-195;
H. Kokubo, T. Tanaka, & Y.O., in Advances in Protein Chemistry and Structural Biology,
T. Karabencheva-Christova (ed.) (Elsevier, Amsterdam, 2013) pp. 63-91.
-
P mu
(E) = n(E)W mu
(E) = const
E
E min
Multicanonical Algorithm
uniform (flat) distribution in energy
W mu
(E) = n(E) -1
Random Walk in Potential Energy Space
MC: B. Berg & T. Neuhaus, Phys. Lett. B267, 249 (1991); Phys. Rev. Lett. 68, 9 (1992).
MD: U. Hansmann, Y.O. & F. Eisenmenger, Chem. Phys. Lett. 259, 321 (1996);
N. Nakajima, H. Nakamura & A. Kidera, J. Phys. Chem. B 101, 817 (1997).
Cf: Wang-Landau method where the weight is dynamically updated
F. Wang & D.P. Landau, Phys. Rev. Lett. 86, 2050 (2001);
Phys. Rev. E 64, 056101 (2001).
-
Canonical Ensemble
MC version:
Multicanonical Ensemble
MC version:
Generalized-Ensemble Algorithms have been
developed in MC algorithms
-
Canonical Ensemble MD version:
MD version:
22
03
mu mui i i i
i
i B
i
E Es sm m m
s E s
sQs s m Nk T Q
s
q q f qq
q
0
1( ) exp( ( ))
( )mu muW E E E
n E
Multicanonical Ensemble
22 3
i i i i
i
i B
i
E s sm m m
s s
sQs s m Nk T Q
s
q q f qq
q
U. Hansmann, Y.O. & F. Eisenmenger, Chem. Phys. Lett. 259, 321 (1996);
N. Nakajima, H. Nakamura & A. Kidera, J. Phys. Chem. B 101, 817 (1997).
-
MULTICANONICAL ALGORITHM
B. Berg & T. Neuhaus, Phys. Lett. B267, 249 (1991).
B. Berg & T. Neuhaus, Phys. Rev. Lett. 68, 9 (1992).
Step 1: Iterations of Short Preliminary Runs to
Determine the Multicanonical Weight Factor Wmu (E)
Step 2: One Long Production Run
Step 3: Analyze the Data to Obtain:
* Global-Minimum Energy Configuration
* Thermodynamic Quantities for Desired Temperatures
(by Ferrenberg-Swendsen Single-Histogram
Reweighting Techniques)
;;
B
C mu
mu
W E TP E T P E
W E
-
30
20
10
0
-10
E
200000150000100000500000
MC Sweeps
Multicanonical
Canonical 50K
Canonical 1000K
-
Single-Histogram Reweighting Techniques
, where ( ) ( .(
)( )
)mu m
u
mu
um N E n E W
N En E
W EE
A. Ferrenberg & R. Swendsen, Phys. Rev. Lett. 61, 2635 (1988).
( ) ; ( )
;
E
C
E ET E
C
E E
n EA E P E T A E e
AP E T en E
Here, the density of states n(E) is obtained from the histogram of the
energy distribution Nmu(E) that was obtained from the production run of
the multicanonical simulation:
-
Enk-ave
-
Single-Histogram Reweighting Techniques
A. Mitsutake, Y. Sugita & Y.O., J. Chem. Phys. 118, 6664 (2003).
When the physical quantity A cannot be written as
a function of E, we use the following equation:
-
fast movie
17-Residue Helical Peptide (120000-300000 MC Sweeps)
Simulation and movie by A. Mitsutake
Canonical MC: T = 200 K Multicanonical MC
-
ST( ; ) exp( ( ))W E T E a T
ST ( )exp( ( )) const( ) dEn E E a TP T
ST( ; ) exp( )m m mW E T E a
exp(am) dEn(E) exp(mE)
am : Dimensionless Helmholtz free energy at temperature Tm
Random Walk in Temperature Space
→ Random Walk in Energy Space
is determinded by iterations of short ST runs am
Discretize Temperature:
A.P. Lyubartsev, et al., J. Chem. Phys. 96, 1776 (1992).
E. Marinari and G. Parisi, Europhys. Lett. 19, 451 (1992).
Temperature is a dynamical variable: Sample temperature uniformly
Simulated Tempering (焼き戻し法)
See also: A. Irback & F. Potthast, J. Chem. Phys. 103, 10298 (1995).
U. Hansmann & Y.O., J. Comput. Chem. 18, 920 (1997).
-
Step 1: Canonical MC/MD Simulations at Temperature Tm
for a Few Steps
Step 2: Temperature is Updated to a Neighboring Value Tm±1
a la Metropolis with Conformations Fixed
where
Repeat These 2 Steps
Canonical Distribution at Any Temperature
by Multiple Histogram Reweighting Techniques (WHAM)
A.P. Lyubartsev, et al., J. Chem. Phys. 96, 1776 (1992).
E. Marinari and G. Parisi, Europhys. Lett. 19, 451 (1992).
Simulated Tempering (焼き戻し法)
-
Multiple-Histogram Reweighting Techniques
(Weighted Histogram Analysis Method: WHAM)
n E
Nm(E)m1
M
nmefm mE
m1
M
, where e
fm n E E
e mE .
A. Ferrenberg & R. Swendsen, Phys. Rev. Lett. 63, 1195 (1989).
S. Kumar, D. Bouzida, R. Swendsen, P. Kollman & J. Rosenberg, J. Comput. Chem. 13, 1011 (1992).
A T
A(E)n E E
e E
n E E
e E
Given M set of histograms Nm(E), which were obtained at Tm, the
following WHAM equations are solved iteratively for density of states n(E)
and dimensionless Helmholtz free energy f m : (nm are the total number
of samples obtained at Tm)
-
A. Mitsutake, Y. Sugita & Y.O., J. Chem. Phys. 118, 6664 (2003).
When the physical quantity A cannot be written as
a function of E, we first obtain the dimensionless
Helmholtz free energy fm (m = 1, …, M) by solving
the WHAM equations. We then use the following
equation:
Multiple-Histogram Reweighting Techniques
(Weighted Histogram Analysis Method: WHAM)
See also: M. R. Shirts and J. D. Chodera, J. Chem. Phys. 129, 124105 (2008).
-
Replica-Exchange Method (also referred to as Parallel Tempering)
1. System
M Non-Interacting Replicas of the Original System at M Different Temperatures
2. Replica-Exchange
Step 1: Independent Canonical Simulations Performed for Each Replica
Step 2: A Pair of Replicas (i and j) Corresponding to Neighboring
Temperatures (Tm and Tn) (i.e., n=m+1) are Exchanged a la Metropolis
Repeat These 2 Steps
3. Canonical Distribution at Any Temperature
by Multiple Histogram Reweighting Techniques (WHAM)
MC: K. Hukushima & K. Nemoto, J. Phys. Soc. Jpn. 65, 1604 (1996).
MD: Y. Sugita & Y.O., Chem. Phys. Lett. 314, 141 (1999).
-
From Multidimensional REM to
Multidimensional MUCA and ST
MMUCA: random walk in multidimensional energy
MST: random walk in multidimensional parameter
MREM: random walk in multidimensional parameter
e.g.,
WHAM eqns.
,
,
,( ), 1
( ) ,
,, 1
( , )
, where ., , m nm n
m n
m n
M
m nE Vm n
ME V EV
m nm n
f
f
N E V
en E V n eE
n
V
e
A. Mitsutake & Y.O., Phys. Rev. E 79, 047701 (2009);
J. Chem. Phys. 130, 214105 (2009); A. Mitsutake, J. Chem. Phys. 131, 094105 (2009).
-
Examples of Multidimensional REM, MUCA, and ST
T. Nagai & Y.O., Phys. Rev. E 86, 056705 (2012).
1. Simulated Tempering and Magnetizing
random walk in temperature T and external field h
*Ising Model
*3-state Potts Model
= h = external field
V = M = magnetization
T. Nagai, Y.O., & W. Janke,
J. Stat. Mech. (2013) P02039.
A. Mitsutake & Y.O., Phys. Rev. E 79, 047701 (2009);
J. Chem. Phys. 130, 214105 (2009); A. Mitsutake, J. Chem. Phys. 131, 094105 (2009).
-
Examples of Multidimensional REM, MUCA, and ST A. Mitsutake & Y.O., Phys. Rev. E 79, 047701 (2009);
J. Chem. Phys. 130, 214105 (2009); A. Mitsutake, J. Chem. Phys. 131, 094105 (2009).
2. Isobaric-Isothermal Ensemble(定圧定温アンサンブル)
* MUCA: Multibaric-Multithermal Algorithm (MUBATH) random walk in potential energy E and volume V H. Okumura & Y.O., Chem. Phys. Lett. 383, 391 (2004). (MC version) H. Okumura & Y.O., Chem. Phys. Lett. 391, 248 (2004). (MD version)
* REM: random walk in temperature T and pressure P Y. Sugita & Y.O., in Lect. Notes in Computational Science & Engineering,
ed. by T. Schlick and H.Gun (2002) pp. 304-332; cond-mat/0102296.
T. Okabe, M. Kawata, Y.O. & M. Mikami, Chem. Phys. Lett. 335, 435 (2001).
Also, see D. Paschek & A. Garcia, Phys. Rev. Lett. 93, 238105 (2004).
* ST: random walk in temperature T and pressure P Y. Mori & Y.O., J. Phys. Soc. Jpn. 79, 074003 (2010).
-
Examples of Multidimensional REM, MUCA, and ST A. Mitsutake & Y.O., Phys. Rev. E 79, 047701 (2009);
J. Chem. Phys. 130, 214105 (2009); A. Mitsutake, J. Chem. Phys. 131, 094105 (2009).
3. Umbrella Sampling
* REM: Replica-Exchange Umbrella Sampling (REUS) random walk in reaction coordinate x Y. Sugita, A. Kitao & Y.O., J. Chem. Phys. 113, 6042 (2000).
* ST: Simulated Tempering Umbrella Sampling (STUS) random walk in reaction coordinate x Y. Mori & Y.O., Phys. Rev. E 87, 023301 (2013). Cf.
* MUCA: Metadynamics (Wang-Landau in reaction coordinate) random walk in reaction coordinate x A. Laio and M. Parrinello, Proc. Natl. Acad. Sci. USA 99, 12562 (2002).
Hk q, p K p E0 q lVl q
l1
L
, where Vl x Kl x q dl 2
.
-
Challenging the prediction of the 3-dimensional structure
of a small protein by MUCAREM.
Villin headpiece subdomain
(36 amino acids; 596 atoms)
sphere of water with radius 30 Å (3513 water molecules);
Total number of atoms = 11,135
Folding of a Small Globular Protein T. Yoda, Y. Sugita & Y.O., Biophys. J. 99, 1637 (2010).
helix1 helix2
MLSDEDFKAVFGMTRSAFANLPLWKQQNLKKEKGLF 1 10 20 30
helix3
Primary Sequence of HP-36
T. Yoda, Y. Sugita & Y.O., Proteins 82, 933-943 (2014).
-
Computational Details
(Force Field = CHARMM22/CMAP for protein
& TIP3P for water)
(1) REMD with 96 replicas in implicit solvent (GB/SA);
initial conformation was fully extended
(2) Unfolded protein w/o any secondary structures was
soaked in a sphere of radius 30Å (with 3513 TIP3P water molecules)
(3) REMD with 128 replicas (T = 250 K ~ 700 K) (4) Determine multicanonical weight factors by WHAM
(iterate several times to refine weight)
(5) Two production runs of MUCAREM with 8 replicas
(MUCAREM1: 1.127 ms in total covering T = 269 K ~ 699 K MUCAREM2: 1.157 ms in total covering T = 289 K ~ 699 K)
T. Yoda, Y. Sugita & Y.O., Biophys. J. 99, 1637 (2010).
-
Villin headpiece subdomain
(36 amino acids; 596 atoms)
in sphere of water of radius 30 Å (3513 water molecules);
altogether 11,135 atoms
MUCAREM simulation
-
MUCAREM2 (Replica 5)
Simulation and movie by T. Yoda
-
Native-Like Structures Obtained from MUCAREM
Main-Chain RMSD = 1.1 Å (residues 2 to 35) [Replica 5] 灰色:自然の構造(PDB ID: 1YRF)、緑色:シミュレーションの結果
T. Yoda, Y. Sugita & Y.O., Biophys. J. 99, 1637 (2010).
-
Challenging the prediction of the 3-dimensional structure
of a small protein by MUCAREM.
Villin headpiece subdomain
(36 amino acids; 596 atoms)
sphere of salted water with radius
30 Å (3494 water molecules,
11 K+, 13 Cl- ≈ 0.2 M KCl);
Salt Effects on Folding of a Small Globular
Protein T. Yoda, Y. Sugita & Y.O., Proteins 82, 933-943 (2014).
helix1 helix2
MLSDEDFKAVFGMTRSAFANLPLWKQQNLKKEKGLF 1 10 20 30
helix3
Primary Sequence of HP-36
-
Native-Like Structure (Global Minimum in Free
Energy) Obtained from MUCAREM Simulation
(Left)
Main-Chain RMSD = 1.25 Å Experimental Structure
(PDB ID: 1YRF)
T. Yoda, Y. Sugita & Y.O., Proteins 82, 933-943 (2014).
-
Free Energy Landscape
in Pure Water in 0.2 M Salted Water
T. Yoda, Y. Sugita & Y.O., Proteins 82, 933-943 (2014).
-
T. Hiroyasu, M. Miki, M. Ogura, & Y. O., J. IPS Japan 43, 70 (2002).
Combinations with Genetic Crossover
Simulated Annealing
Y. Sakae, T. Hiroyasu, M. Miki, K. Ishii & Y. O., J. Phys.: Conf. Ser. 487,
012003 (2014).
Y. Sakae, T. Hiroyasu, M. Miki, K. Ishii & Y. O., in preparation.
Metropolis
Y. Sakae, T. Hiroyasu, M. Miki & Y. O., J. Comput. Chem. 32, 1353 (2011).
-
High T A B C D F E
GA crossover
GA crossover
GA crossover
GA crossover
Low T
Y. Sakae, T. Hiroyasu, M. Miki & Y. O., J. Comput. Chem. 32, 1353 (2011).
Parallel Simulated Annealing MD with Genetic Crossover
(PSAMD/GAc2)
Step 1. Parallel simulated annealing simulations for a certain MD steps
Step 2. Genetic crossover Repeat these two steps. P
aral
lel
Sim
ula
ted A
nnea
lin
g S
imula
tions
-
All dihedral angles in randomly selected
consecutive amino-acid residues
are exchanged.
Structure A
Structure B
Dihedral Angle
Exchange a Randomly Chosen Pair of Dihedral Angle Sets
Genetic Crossover (2-Point Crossover)
-
1. All dihedral angles in randomly selected n (2-10)
consecutive amino-acid residues are exchanged.
2. A short (say, 20 ps) MD simulation with
H = H0 + Hconstr where Hconstr is a harmonic constraint potential that
constrains the corresponding main-chain dihedral
angles (f, y).
3. A short (say, 20 ps) equilibration MD with
H = H0 4. Selection rule is imposed with respect to the final
conformations in Step 3.
After Genetic Crossover Operation,
two children will have large energies;
side chains bump into each other.
Y. Sakae, T. Hiroyasu, M. Miki, K. Ishii & Y. O., J. Phys.: Conf. Ser. 487,
012003 (2014).
-
Detailed Balance Conditions
1. All dihedral angles in randomly selected n (2-10)
consecutive amino-acid residues are exchanged.
2. A short (say, 20 ps) MD simulation with
H = H0 + Hconstr where Hconstr is a harmonic constraint potential that
constrains the corresponding main-chain dihedral
angles (f, y).
3. A short (say, 20 ps) equilibration MD with
H = H0 4. Accept or reject a la Metropolis criterion.
Detailed Balance Condition is satisfied just as in
Hybrid Monte Carlo method, provided that we use a
volume-preserving and time-reversal MD integrator.
Metropolis with Genetic Crossover Combined
-
Example 1
Trp-Cage (PDB ID: 1L2Y)
20 residues
method: MD (Langevin dynamics)
temperature: 282 K
solvent: GB/SA
force field: AMBER ff03
no. of individuals: 16
simulation time per individual: 100ps×100(10ns)
Simulation(Individual No.2) PDB Structure(NMR)
RMSD : 0.809 Å
Y. Sakae, T. Hiroyasu, M. Miki, K. Ishi & Y.O., in preparation.
-
Example 2
Villin Headpiece (PDB ID: 1YRF)
36 residues
PDB Structure (X-ray)
RMSD : 2.234 Å
Simulation(Individual No. 11)
method: MD (Langevin dynamics)
temperature: 300 K
solvent: GB/SA
force field: AMBER ff03
no. of individuals: 32
simulation time per individual: 200ps×100(39.4ns)
Y. Sakae, T. Hiroyasu, M. Miki, K. Ishi & Y.O., in preparation.
-
Example 3
Protein A (PDB ID: 1BDD)
46 residues (10-55 out of 60)
RMSD : 1.707 Å Simulation(Individual No. 5)
PDB Structure(NMR)
method: MD (Langevin dynamics)
temperature: 300 K
solvent: GB/SA
force field: AMBER ff03
no. of individuals: 32
simulation time per individual: 1.0ns×90(90ns)
Y. Sakae, T. Hiroyasu, M. Miki, K. Ishii & Y. O., J. Phys.: Conf. Ser. 487,
012003 (2014).
-
2-Dimensional ST Simulation
in Isobaric-Isothermal Ensemble
temperature and pressure become dynamical
variables.
2-dimensional random walk
in temperature and pressure
Y. Mori & Y.O., J. Phys. Soc. Jpn. 79, 074003 (2010);
in preparation.
-
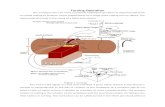
Pressure-Induced Unfolding of Ubiquitin
R. Kitahara et al. (2005)
30 bar – 3000 bar
NMR Experiments
• 76 amino acids
• 6232 water molecules
• 19985 atoms
PDB: 1V80
Simulated System
-
Time series of pressure P, potential energy E, and volume V for ubiquitin
Y. Mori & Y.O., in preparation.
-
Large structural fluctuations
Amino Acid Residues
f [Å]
• Fluctuations of distance d between pairs of Ca atoms.
Large fluctuations observed in agreement with experiments.
http://maru.bonyari.jp/texclip/texclip.php?s=/begin{align*}f /equiv /sqrt{/langle d^2 /rangle - /langle d /rangle^2}/end{align*}
-
Structural changes under high pressure
r
r [Å]
distribution [Å-1]
r
-
Ubiquitin and water molecules
at low pressure at high pressure
Simulation and movie by Y. Mori
-
Experiments:
R. Kitahara & K. Akasaka,
PNAS 100, 3167 (2003).
N
H
-
Calculation of chemical shifts • We calculated 15N chemical shifts for all the amino acid residues
and show the distributions of several calculated chemical shifts.
• Program: CamShift (ver. 1.35)
• Pressure: 1bar (blue) to 4,000 bar (red)
Residue 70
Residue 68 Residue 69
Residue 71 Residue 72
low pressure
high pressure
Y. Mori et al., in preparation.
-
Prediction of Protein-Ligand Binding Structures by
Replica-Exchange Umbrella Sampling H. Kokubo, T. Tanaka & Y.O., J. Comput. Chem. 32, 2810-2821 (2011).
Replica-Exchange Umbrella Sampling (REUS)
Potential of Mean Force
Hk q, p K p E0 q lVl q
l1
L
, where Vl x Kl x q dl 2
.
W x 1
ln P ,0 x .
P , E,x Nm(E,x)e
EV x
m1
M
nmef m m EVm x
m1
M
,
e f m P
m , mE,x
x
E
.WHAM
Y. Sugita, A. Kitao & Y.O., J. Chem. Phys. 113, 6042 (2000).
See also:
E.M. Boczko & C.L. Brooks, J. Phys. Chem. 97, 4509 (1993).
B. Roux, Comp. Phys. Commun. 91, 275 (1995).
-
Five test systems [ligand (protein)] (T = 300 K, P = 1 atm)
benzodiazepine
(protein: MDM2)
deoxythymidine 3’,5’-bisphosphate (pdTp)
(protein: staphylococcal nuclease)
tyrosine
(protein: aldolase)
cytidylic acid (2’-CMP)
(protein: ribonuclease A)
2-aminopyrimidine
(protein: heat shock protein HSP90)
PDB code
PDB code
H. Kokubo, T. Tanaka & Y.O., J. Comput. Chem. 32, 2810-2821 (2011).
-
Umbrella Potentials (24 Replicas)
dm : 5.0, 5.5, 6.0, 6.5, 7.0, 7.5, 8.0, 8.5, 9.0, 9.5, 10.0, 10.5, 11.0,
12.0, 13.0, 14.0, 15.0, 16.0, 17.5, 19.0, 20.5, 22.0, 23.5, 25.0
km : 1.0 for dm < 13.5 Å, 0.5 for dm > 13.5 Å
MD simulation: 110-220 nsec per replica
H. Kokubo, T. Tanaka & Y.O., J. Comput. Chem. 32, 2810-2821 (2011).
-
The initial structures of five protein-ligand complexes
53
The space-filled molecules, which do not actually exist in these
simulations, show the correct ligand binding positions (from
PDB) as references. The ligands are in bulk water and far
away from the binding pockets. 53
-
Simulation and movie by H. Kokubo
MDM2 and benzodiazepine: REUS simulation
-
Snapshots of MDM2 protein system
Protein surface is fluctuating
-
Simulation and movie
by H. Kokubo
heat shock protein and
2-aminopyrimidine
Asp-78 and ligand
-
Results: REUS with 24 replicas
• Starting from configurations in which protein and ligand are far away from
each other in each system, our method predicted the ligand binding
structures as the global minima in free energy (or, potential of mean force)
in excellent agreement with the experimental data from PDB.
potential of mean force shows
the most stable distance
crystal
predicted
We remark that for 1ROB and 1SNC, there
are attempts by a popular existing docking
program, GOLD, but they failed in the
predictions (classified as significant errors).
H. Kokubo, T. Tanaka & Y.O., J. Comput. Chem. 32, 2810-2821 (2011).
-
Kinase systems we tested
JNK3
1PMV 150 nM
2O2U 3.0 uM
P38
1OVE 0.74nM
1OZ1 6.5 nM
*240ns REUS simulations were performed for four systems:
1PMV, 2O2U, 1OVE, and 1OZ1.
H. Kokubo, T. Tanaka & Y.O., J. Chem. Theor. Comput. 9, 4660-4671 (2013).
-
Potential of Mean Force
59
1OZ1 1OVE
1PMV 2O2U
150 nM 3000 nM
0.74nM 6.5 nM
p38 p38
JNK3 JNK3
H. Kokubo, T. Tanaka & Y.O., J. Chem. Theor. Comput. 9, 4660-4671 (2013).
-
The comparison of the predicted global minimum free
energy structures and PDB structures
1PMV
1OVE
2O2U
JNK3 JNK3
p38 p38
1OZ1 crystal predicted
H. Kokubo, T. Tanaka & Y.O., J. Chem. Theor. Comput. 9, 4660-4671 (2013).
-
Movie for 1PMV
Simulation and movie by H. Kokubo
-
Necessity of protein flexibility
P38 & dihydroquinolinone (PDB ID: 1OVE)
Without flexibility With flexibility
H. Kokubo, T. Tanaka & Y.O., J. Chem. Theor. Comput. 9, 4660-4671 (2013).
-
Necessity of protein flexibility
P38 & dihydroquinolinone (PDB ID: 1OVE)
Without flexibility With flexibility
H. Kokubo, T. Tanaka & Y.O., J. Chem. Theor. Comput. 9, 4660-4671 (2013).
-
Prediction of Protein-Ligand Binding Structures by
2-dim H-REMD: Replica-Exchange Umbrella
Sampling (REUS) and Replica-Exchange Solute
Tempering (REST)
H. Kokubo, T. Tanaka & Y.O., J. Comput. Chem. 34, 2601-2614 (2013) .
Potential Energy:
[ ] [ ] [ ]] [
0
[ ]
0
( ) ( ) () )( ( )ni i i imn ll ls ss mn iE q U q U qU Vq q
l: ligand
s: protein/water
REST: P. Liu, B. Kim, R.A. Friesner, & B.J. Berne, PNAS 102, 13749 (2005).
REUS: Y. Sugita, A. Kitao & Y.O., J. Chem. Phys. 113, 6042 (2000).
Total no. of replicas: M × N REST parameters (n=1, 2, …, N)
REUS parameters, i.e., umbrella potentials
(m=1, 2, …, M)
-
2-dim H-REMD: REUS (M=24) + REST (N=8)
(192 replicas)
REUS (1T4E) REUS/REST (1T4E)
No. of
Random Walk
Cycles
H. Kokubo, T. Tanaka & Y.O., J. Comput. Chem. 34, 2601-2614 (2013) .
-
Force field refinement for all-atom
protein models
with Yoshitake Sakae
REVIEW:
Y. Sakae & Y.O.,
in Computational Methods to Study the Structure and Dynamics of Biomolecules and
Biomolecular Processes – from Bioinformatics to Molecular Quantum Mechanics
A. Liwo (ed.) (Springer-Verlag, Berlin Heidelberg, 2014) pp. 195-247.
-
Force-field parameters for all-atom models
2
2
12 6
( )
( )
[1 cos( )]2
332
r eq
eq
n
ij ij i
conf
bonds
angles
dihedrals
i j j
j
i ij ij
K r
K
E r
Vn
A q q
R R
B
R
f
Bond-stretching
Bond-bending
Dihedral angle
Non-bonding interactions
(Lennard-Jones and electrostatic)
These energy terms include some force-field parameters (blue color)
Commonly used conformational potential energy
The existing force fields have different force-field parameters
-
Force-field dependency of secondary-structure properties
C-peptide
(13 residues)
AMBER parm94
AMBER parm96
AMBER parm99
CHARMM22
OPLS-AA/L
GROMOS96
α-h
elix
T. Yoda, Y. Sugita & Y.O.,
Chem. Phys. Lett. 386, 460 (2004).
Helicity of C-peptide
-
Typical example of folding simulations using different force fields
AMBER ff94 AMBER ff96
C-peptide
(13 residues) Lys-Glu-Thr-Ala-Ala-Ala-Lys-Phe-Glu-Arg-Gln-His-Met
Method:Simulated annealing, Simulation time : 1.0 nsec, Temperature : 700~200 K, Solvent model : GB/SA
-
70
Conformational Energy
nonbondtorsionBABLconf EEEEE
rest
n
nn
torsion
EE
nV
E
),(
)]cos(1[2
y
Backbone-torsion energy term
backbone dihedral
angles φ and ψ
rest of the torsion terms
Φ : all dihedral angles
n : number of waves
γn : phase
Vn : Fourier coefficient
Side chain
-
Backbone-torsion energy surfaces of some force fields
-180
-90
0
90
180
180
90
0
-90
-180f
y
-180
-90
0
90
180
180
90
0
-90
-180f
y-180
-90
0
90
180
180
90
0
-90
-180f
y
-180
-90
0
90
180
180
90
0
-90
-180f
y -180
-90
0
90
180
180
90
0
-90
-180f
y-180
-90
0
90
180
180
90
0
-90
-180f
y
AMBER parm94 AMBER parm96 AMBER parm99
CHARMM22 OPLS-AA OPLS-AA/L
-
72
m n
nn
mm n
Vm
VE )]cos(1[
2)]cos(1[
2),( yy
resttorsion EE ),( y
)sinsincossin yy nminnh mnmn
1 1
11
sincoscoscos(
)sincos()sincos(),(
m n
mnmn
n
nn
m
mm
nmgnmf
nendmcmba
yy
yyy
Y. Sakae and Y.O., J. Phys. Soc. Jpn. 75, 054802 (9 pages) (2006).
1. Proposal of new functional form
conventional energy term
New torsion energy term
New backbone-torsion-energy term
a,bm,cm,dn,en,fmn,gmn,hmn,imn
: Fourier coefficient
-
Ramachandran plot
タンパク質の構造入門第2版
Example of the application of new backbone-torsion energy term
-180
-90
0
90
180
180
90
0
-90
-180f
y-180
-90
0
90
180
180
90
0
-90
-180f
y-180
-90
0
90
180
180
90
0
-90
-180f
y
a-helix region -structure region
Energy surface of new energy term can represent Ramachandran space directly
-
74
-180
-90
0
90
180
180
90
0
-90
-180f
y
-180
-90
0
90
180
180
90
0
-90
-180f
y
~
~
~
~
~
~
~
~
AMBER parm94
AMBER parm96
Application to AMBER parm94 and AMBER parm96
αhelix βstructure
αhelix βstructure
-
75
AMBER parm94 AMBER parm96
Original Original
a-helix region a-helix region
-structure region -structure region
Results of folding simulations C-peptide
Simulated annealing simulation Simulation time : 1ns (1,000,000 MD steps × 1.0fs × 60 times) Temperature : 2,000K to 250K (Berendsen’s method)
Solvent : GB/SA model
-
76
AMBER parm94 AMBER parm96
G-peptide
Simulation time : 1ns (1,000,000 MD steps × 1.0fs × 60 times) Temperature : 2,000K to 250K (Berendsen’s method)
Solvent : GB/SA model
Results of folding simulations
Original Original
a-helix region a-helix region
-structure region -structure region
Simulated annealing simulation
-
mN
mif
2
1 1
1 m
m
m
NN
i
m im
F fN
m
m
i
m
toti
x
Ef
}{
}{m
totEmi
fAtom i
Molecule m
Optimization method of force-field parameters
Y. Sakae and Y.O., Chem. Phys. Lett. 382, 626-636 (2003)
Number of atoms in molecule m
Force acting on atom i
Total potential energy for molecule m
If force-field parameters are of ideal values, all native structures
are stable without any force acting on each atom in molecules.
(we expect F=0)
In reality, F≠0, and because F ≥0, we can optimize the force-field
parameters by minimizing F with respect to these parameters.
-
Structures of selected 100 proteins
Resolution: >= 2.0Å, Sequence similarity of amino acid: >= 30%, Number of residue: < 200
-
Results: Optimized force-field parameters and its energy surface
-180
-90
0
90
180
180
90
0
-90
-180f
y-180
-90
0
90
180
180
90
0
-90
-180f
y
AMBER parm94 AMBER parm96
Optimized ff
yy
yy
yy
yy
y
sinsin603.0cossin114.0
sincos247.0coscos427.0
2sin054.02cos019.0
sin160.0cos287.0
2sin100.02cos088.0
sin327.0cos835.0),(
-180
-90
0
90
180
180
90
0
-90
-180f
y
-
80
C-peptide
(13 residues)
blocked by COCH3- and –NH2
G-peptide
(16 residues)
Test simulations
Y. Sugita and Y.O., Chem. Phys. Lett. 314, 141-151 (1999)
Replica Exchange Molecular Dynamics (REMD) simulation
Simulation time : 5.0 ns (32 replica)
Temperature : 700K to 250K (Nosé-Hoover method)
Solvent : GB/SA model
Program : Modified TINKER program package
-
C-peptide
300K
Comparison of helicity and strandness
-
G-peptide
300K
Comparison of helicity and strandness
-
C-peptide
),(ln),( yfyf BB PTkG
Optimized ff
AMBER ff96 AMBER ff94 300K
Potential of mean force
-
G-peptide
),(ln),( yfyf BB PTkG
Optimized ff
AMBER ff96 AMBER ff94 300K
Potential of mean force
-
COLLABORATORS Ayori MITSUTAKE [IMS Keio Univ.]
Yuji SUGITA [IMS Univ. Tokyo RIKEN]
Takao YODA [IMS Nagahama Inst. Bio-Science]
Yoshitake SAKAE [IMS Hiroshima Univ. Nagoya Univ.]
Yoshiharu MORI [Nagoya Univ. IMS]
Tetsuro NAGAI [Nagoya Univ. Ritsumeikan Univ.]
Hironori KOKUBO [IMS Univ. of Houston Takeda Pharm.]
Toshimasa TANAKA [Takeda Pharm.]
Takeshi NISHIKAWA [IMS AIST FOCUS]
Yasuyuki ISHIKAWA [Univ. Puerto Rico]
Ryo KITAHARA [Ritsumeikan Univ.]
Kazuyuki AKASAKA [Kinki Univ.]
Wolfhard JANKE [Univ. Leipzig]
Giovanni LA PENNA [ICCOM, CNR, Firenze]
Michele VENDRUSCOLO [Univ. of Cambridge]
Christopher M. DOBSON [Univ. of Cambridge]