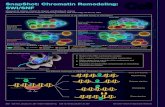
Nucleosome Positioning
description
Transcript of Nucleosome Positioning

Nucleosome Positioning
Histones and DNA Bending

DNA packaging
• 3 X 109 base pairs in human genome• ~1 m if unraveled• Compacted into nucleus
– 100 m in diamter

Histones

Histones

PDBID = 1KX3

Nucleosome Arrays
Dorigo B, et al. Science. 2004 306(5701):1571-3.

Current Opinion in Structural Biology 2006, 16:336–343

http://www2.kenyon.edu/Depts/BioEllipse/courses/biol114/Chap01/chrom1.gif

Histones vs. DNA
• Histones fold DNA• DNA sequence determines histone
binding– Does histone bind sequence?
• Absence of direct sequence specific recognition
– Bending flexibility of DNA determines affinity for complementary protein surface

DNA as a Rod

Twist
Bend


Flexibility
• Torsional Flexibility– Angle between base pairs
• Bending Flexibility– Properties of individual base steps
• Flexibility of DNA is a Sequence-dependent property

Flexibility determinants
• Hydrogen bonds• Electronic configuration of base pairs• Base Pair Stacking energy• Exocyclic groups in major and minor
grooves

Base Pairing
http://en.wikipedia.org/wiki/Base_pair

Base Pairing

Base Stacking
http://researchnews.osu.edu/archive/dna_base_stacking.jpg

Base Stacking
Richmond & Davey 2003

Electrostatics
Philos. Transact. A Math Phys. Eng. Sci. 2004 Jul 15;362(1820):1423-38.


Sequence-directed positioning
Widlund et al. 1997• Mouse Genome• Micrococcal nuclease• 146 bp DNA fragments• DNA seq added to end• Amplify using PCR• 10 rounds cycles
– competitive nucleosome assembly
Chromatin: Structure and Function Alan P. Wolffe

Intrinsically curved DNA sequences
• Oligo(dA).oligo(dT) tracts (4-6bp)– Phased with a periodicity similar to that
of DNA helix
• TATA boxes repeated every 10 bp• CA and CTG di/tri-nucleotide repeats

Energy Landscape

Matrix Method
• Nucleotide phasing of 10bp repeats influences bending of DNA

Periodic occurrence of A/T & GC
• A/T & G/C out of phase
• A/T narrow minor groove
• G/C wide minor groove
Satchwell et al. J. Mol. Biol. 1986 191:659-75





Minor Groove bending facilitated by Arg side chain into minor groove when it faces histone octamer
Major groove narrows to ~9Å
Minor groove narrows to ~3Å

Structure of DNA in Nucleosome
• Discontinuous curvature
Richmond & Davey 2003

Summary
Ong, Richmond & Davey 2007
• Major Gray• Minor White

Unusual DNA conformation parameters induced by histone
binding• Implicates sequence-dependent
– protein recognition– nucleosome positioning– nucleosome mobility
• Can we predict DNA:Nucleosome binding based on sequence?












![Computational biology: deep learning...from DNA sequence, RNA polymerase binding, nucleosome positioning and transcriptional data [16], as well as gene expression from histone modifications](https://static.fdocuments.net/doc/165x107/61487fc62918e2056c22ba9d/computational-biology-deep-learning-from-dna-sequence-rna-polymerase-binding.jpg)