MASS SPECTROMETRY & STANFORD CHEMISTRY · 4 The Mass Spectrometer: Components 1.Ion...
Transcript of MASS SPECTROMETRY & STANFORD CHEMISTRY · 4 The Mass Spectrometer: Components 1.Ion...

MASS SPECTROMETRY & MASS SPECTROMETRY & STANFORD CHEMISTRYSTANFORD CHEMISTRY
Allis S. Chien, Ph.D.Friday November 21, 2003

Brief intro to Mass SpectrometersIonization: ESI & APCISUMS InstrumentationMS – Organic, Organometallic &
BiomoleculeHPLC-MS
MASS SPECTROMETRY & MASS SPECTROMETRY & STANFORD CHEMISTRYSTANFORD CHEMISTRY

3
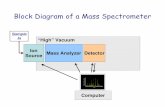
The Mass Spectrometer: Operation
Steps in generating a mass spectrum:
1. Produce ions2. Separate or filter ions3. Detect ions4. Process the data

4
The Mass Spectrometer: Components
1.Ion source/interface2.Mass analyzer, including:
a.Mass analyzer (quadrupole, ion trap, TOF, etc.)b.Vacuum systemc.Some electronics
3.Detector4.Data storage, (processing), and output device
(usually a computer)

5
Quadrupole Ion Trap
LC Pump ESI Quadrupole Ion Trap
Syringe Pump Detector

6
What is API?Atmospheric Pressure Ionization
ESI – Electrospray IonizationSolution-phase process (for the most part)
APCI – Atmospheric Pressure Chemical Ionization
Gas-phase process
An interface between HPLC and Mass Detection
Designed to separate and ionize analytes from HPLC solvents

7
ESI Needle+/- 5 kV
Heated Capillary
Taylor Cone
Solvent evaporation and ion release
+ ++++ +
+ +
+ ++++ +
+ +
+
++
+++++ +
++
+ ++++ +
+ +
+ ++++ +
+ +
+ +++
++
++
++
+
++ ++ +
+ +
+++
+ +
++ ++ +
+++
+ +
+
+++
+++
+
+
+
++
+
++
+
++
++
++
+
+
+
+
+
Electrospray – Basic Layout

APCI: Atmospheric Pressure Chemical Ionization
Mechanism for positive ion formationPrimary ion formation:
Secondary ion formation:
Analyte ion formation:
−•+−
−•+−
+→+
+→+
ee
ee
2OHOH
2NN
22
22
OHOHOHOH 322•+•+ +→+
OH]H[OH 23 ++→+ ++ AnalyteAnalyte

9
ESI or APCI ?
Many compounds can be analyzed by both techniques with different sensitivities
ESI is for highly polar compounds
ESI is for molecular weights >1000 amu
ESI is for thermally fragile compounds
APCI generally gives more fragmentation

Analyte Compatibility
ESIESI
EIEI PBIPBITSPTSP FABFAB
Mol
ecul
ar
Wei
ght
200,000
15,000
1,000
Non Polar
APCI
Polar

SUMS Instrumentation

12
LCQ Classic MS
• Quadrupole Ion Trap LC-MS• ThermoFinnigan Surveyor HPLC &
LCQ “Classic” MS
– MW determination– Analytical LC-MS– MSn

13
LCQ Deca XP Plus MS
– Capillary LC-MS– Protein identification & characterization– Complex protein mixture analysis– Assays, quantitation
•LC Packings Capillary HPLC System & ThermoFinnigan LCQ Deca XP Plus MS

14
Q-Tof API
MicromassQ-Tof
•High resolution MS•Protein identification & characterization•De novo peptide sequencing•Post-translational modification ID
•Hybrid Tandem Quadrupole – Time of Flight MS

Mass Spec Examples
Organic CompoundsOrganometallic CompoundsBiomolecules
LC-MSHigh Resolution MS

16
Commonly Observed Ions in ESI
NH
O
OH
NH
OHO
O
NH
O
O
O
NH
C27H42N4O8
MW 550.3
100
0
20
40
60
80
Rel
ativ
e Ab
unda
nce
573.1
574.1
551.0[M+H]+
[M+Na]+
Hiroko Tanaka
+ESI

17
Commonly Observed Ions in ESI
NH
O
OH
NH
OHO
O
NH
O
O
O
NH
C27H42N4O8
MW 550.3
540 550 560 570 580m/z
0.4
0.0
0.1
0.2
0.3
100
0
20
40
60
80
Rel
ativ
e Ab
unda
nce
573.1
574.1
551.0
549.0
550.0 571.0
[M+H]+
[M+Na]+
[M+Na-2H]-
[M-H]-
Hiroko Tanaka
+ESI
-ESI

18
N
O
NH
O
O
O
O
OH
OO
N O
O
O
FF F
FF
N3
98
0
20
40
60
80
Rel
ativ
e Ab
unda
nce
1007.2
951.21023.1677.2 855.3618.3
C48H53N6O11F5
MW 984.4Hiroko Tanaka
Na+ Adduct MS

19
N
O
NH
O
O
O
O
OH
OO
N O
O
O
FF F
FF
N3
400 600 800 1000m/z
100
0
20
40
60
80
98
0
20
40
60
80
Rel
ativ
e Ab
unda
nce
1007.2
951.21023.1677.2 855.3618.3
951.1
1007.1
732.1618.1 923.1
534.1C48H53N6O11F5
MW 984.4Hiroko Tanaka
MS/MS Does Not Displace Na+
MS/MS
MS

20
APCI of a Non-polar Compound
NN
O
O
C52H74N2O2
MW 758.6
200 400 600 800 1000 1200 1400m/z
0
10
20
30
40
50
60
70
80
90
100
Rel
ativ
e Ab
unda
nce
759.7
Ned Bowden
[M+H]+

21
APCI of a Non-polar Compound
NN
O
O
C52H74N2O2
MW 758.6
200 400 600 800 1000 1200 1400m/z
759.7
Ned Bowden
8 760 762m/z
759.7
760.7
761.7
[M+H]+

22
APCI of a Non-polar Compound
NN
O
O
C52H74N2O2
MW 758.6Average MW 759.2
200 400 600 800 1000 1200 1400m/z
759.7
Ned Bowden
8 760 762m/z
759.7
760.7
761.7
[M+H]+
759.7

23
Cu Isotope PatternSimulat ionCuProf ileResolut ion:
Daltons 1at 5% height
Charges 1Chrg d ist 0Ions 2M in Ion Ab. 1e-020M in Ions 5000M ax Ions. 20000
61 62 63 64 65 66 67m/z
0.00
0.05
0.10
0.15
0.20
0.25
0.30
0.35
0.40
0.45
0.50
0.55
0.60
0.65
Abun
danc
e
62 .9
64.9

24
Cu Compound: Theoretical MS
N NCu
O O
H H
C36H56N2O2CuMW 611.37
Russell Pratt
C36 H56 N2 O2 Cu1Simulat ionC36H56N2O2CuProf ileResolut ion:
Daltons 1at 5% height
Charges 1Chrg d ist 0Ions 3328M in Ion Ab. 1e-020M in Ions 5000M ax Ions. 20000
610 612 614 616m/z
0.00
0.05
0.10
0.15
0.20
0.25
0.30
0.35
0.40
0.45
Abun
danc
e
611.4
613.3
612.4
614.4
615.4616.4 617.4

25
Cu Compound: +ESI and -ESI
608 610 612 614 616m/z
6
0
2
4
100
0
20
40
60
80
Rel
ativ
e Ab
unda
nce
612.3
614.3
615.3
610.3
612.3611.2
N NCu
O O
H H
C36H56N2O2CuMW 611.37
Russell Pratt
+ESI
-ESI

26
Zr Isotope PatternSimulat ionZrProf ileResolut ion:
Daltons 0.35at 5% height
Charges 1Chrg d ist 0Ions 5M in Ion Ab. 1e-020M in Ions 5000M ax Ions. 20000
88 89 90 91 92 93 94 95 96 97m/z
0.00
0.05
0.10
0.15
0.20
0.25
0.30
0.35
0.40
0.45
0.50
Abun
danc
e
89 .9
93.991.9
90.9
95.9

27
Zr Compound: Theoretical MS
ZrO
O
N
N
C28H46N2O2ZrMW 532.26
Kuo-Wei Huang
532 533 534 535 536 537 538 539m /z
533.26
534.26
535.26
537.26
536.27 538.27539.26
Simulated [M+H]+

28
Theoretical vs. Experimental
ZrO
O
N
N
C28H46N2O2ZrMW 532.26
Kuo-Wei Huang
532 533 534 535 536 537 538 539m /z
533.26
534.26
535.26
537.26
536.27 538.27539.26
532 533 534 535 536 537 538 539 540
533.1
534.1535.1
537.1
536.1 538.1 539.1
Simulated [M+H]+
Experimental [M+H]+

29
Loss of Counterion
500 600 700 800 900 1000 1100 1200m/z
849.3
Richard Decreau
C37H8F15N4FeClMW 884.0
N
N N
NFF
F
F F F F
F F
FFF
F
FF
Fe
Cl

30
Loss of Counterion
500 600 700 800 900 1000 1100 1200m/z
849.3
Richard Decreau
[C37H8F15N4Fe]+
MW 849.0
C37H8F15N4FeClMW 884.0
N
N N
NFF
F
F F F F
F F
FFF
F
FF
Fe
Cl[M-Cl]+

31
Loss of Counterion
500 600 700 800 900 1000 1100 1200m/z
849.3
45 850m/z
849.3
850.3
851.3847.3
Richard Decreau
[C37H8F15N4Fe]+
MW 849.0
C37H8F15N4FeClMW 884.0
N
N N
NFF
F
F F F F
F F
FFF
F
FF
Fe
Cl[M-Cl]+

32
FePd-Porphyrin
1400 1500 1600 1700 1800 1900 2000m/z
0
10
20
30
40
50
60
70
80
90
100
Rel
ativ
e Ab
unda
nce
1558.0
1601.9
Richard Decreau
NN
NN
HN
NHFe
O
N N ON
NNH
NNO
N
F3C
MeMeMe
Pd
HNN
O
ClCl
C77H55O4N16F3FePdCl2MW 1558.0

33
[M]+
0 1600 1700 1800 1900 2000m/z
1558.0
1601.9
Richard Decreau
NN
NN
HN
NHFe
O
N N ON
NNH
NNO
N
F3C
MeMeMe
Pd
HNN
O
ClCl
C77H55O4N16F3FePdCl2MW 1558.0
155 2 15 5 4 15 5 6 15 58 15 6 0 15 6 2 15 6 4 15 6 6m / z
15 58 .0
155 6 .1
15 57 .1 15 6 0 .0
15 55 .0 15 6 1.0
15 6 1.9
15 5 4 .1 15 6 3 .0
15 6 4 .115 5 3 .0

34
Fe(II) Oxidized to Fe(III)
0 1600 1700 1800 1900 2000m/z
1558.0
1601.9
Richard Decreau
NN
NN
HN
NHFe
O
N N ON
NNH
NNO
N
F3C
MeMeMe
Pd
HNN
O
ClCl
C77H55O4N16F3FePdCl2MW 1558.0
155 2 15 5 4 15 5 6 15 58 15 6 0 15 6 2 15 6 4 15 6 6m / z
15 58 .0
155 6 .1
15 57 .1 15 6 0 .0
15 55 .0 15 6 1.0
15 6 1.9
15 5 4 .1 15 6 3 .0
15 6 4 .115 5 3 .0
15 5 5 15 6 0 15 6 5m / z
15 5 8 . 2 0
15 5 6 . 2 015 5 9 . 2 0
15 6 1. 2 015 5 5 . 2 0
15 6 2 . 2 0
15 5 4 . 2 0 15 6 3 . 2 015 6 4 . 2 015 5 3 . 2 0 15
theoretical

35
Exchangeable Hydrogens
220 222 224 226m /z
NH NH
NH
C13H21N3
MW 219.2
Xavi Ribas

36
H/D Exchange Over Time
220 222 224 226m /z
223.3
C13H21N3MW 219.2C13H18D3N3
MW 222.2
Xavi Ribas
1 min
2 min
3 min
4 min
6 min
ND DN
ND

37
H2N
NH
N
N
O
NH2N
O
O
PO
O
OH
NH
N
N
O
NH2N
O
O
PO
O
OH
NH
N
N
O
NH2N
O
O
PO
O
OH
O
O
PO
O
OH
NH
N
N
O
NH2N
O
OH
NH
N
N
O
NH2N
Oligonucleotides: -ESI
800 1000 1200 1400 1600m/z
0
10
20
30
40
50
60
70
80
90
100
Rel
ativ
e Ab
unda
nce
1581
.6
790.
580
1.4
1603
.616
25.6
C50H62N26O27P4
MW 1582.3
Greg Miller
GGGGG
-ESI

38
Doubly Charged Ion [M-2H]2-
790 795 800 805 810 815m/z
0
10
20
30
40
50
60
70
80
90
100
Rel
ativ
e Ab
unda
nce
790.
579
0.9
801.
480
1.9
802.
2
812.
381
2.7
H2N
NH
N
N
O
NH2N
O
O
PO
O
OH
NH
N
N
O
NH2N
O
O
PO
O
OH
NH
N
N
O
NH2N
O
O
PO
O
OH
O
O
PO
O
OH
NH
N
N
O
NH2N
O
OH
NH
N
N
O
NH2N
C50H62N26O27P4
MW 1582.3
Greg Miller

39
Peptide Charge States
400 600 800 1000 1200 1400 1600m/z
0
10
20
30
40
50
60
70
80
90
100
Rel
ativ
e Ab
unda
nce
819.2
1228.2
710.1 1050.2615.2

40
Peptide MW = 2455 Da
400 600 800 1000 1200 1400 1600m/z
0
10
20
30
40
50
60
70
80
90
100
Rel
ativ
e Ab
unda
nce
819.2
1228.2
710.1 1050.2615.2
[M+2H]2+
[M+3H]3+
[M+4H]4+
Neutral mass = (819.2 * 3)-3

41
Proteins: Multiple charge states031009_12216' # 1042-1147 RT: 27.33-30.13 A V: 106 NL: 2.73E7T: + p ESI Full ms [ 400.00-2000.00]
600 800 1000 1200 1400 1600 1800 2000m/z
0
10
20
30
40
50
60
70
80
90
100
Rel
ativ
e A
bund
ance
948.0903.1
972.5
997.9842.9 1115.1
1148.91263.8 1353.9824.8
774.2 1404.0 1579.3743.9
1648.2729.7 1723.1
1806.3 1893.3690.5 1995.1
Ian Suydam

42
Deconvoluted Protein Mass
20000 25000 30000 35000 40000 45000 50000 55000mass
0
10
20
30
40
50
60
70
80
90
100
Rel
ativ
e A
bund
ance
37884.0
50629.0 54668.047473.041979.025315.0 33798.028480.020990.0
Ian Suydam

43
Deconvoluted Protein Mass
20000 25000 30000 35000 40000 45000 50000 55000mass
0
10
20
30
40
50
60
70
80
90
100
Rel
ativ
e A
bund
ance
37884.0
50629.0 54668.047473.041979.025315.0 33798.028480.020990.0
Ian Suydam
38000mass
37884.0
38059.0
37560.0 38632.0

44
Charge State Series
700 800 900 1000 1100 1200 1300 1400m/z
+261458.2
+271404.0
+281353.9+29
1307.3
+301263.8
+311223.0
+321184.7
+331148.9
+351083.5
+361053.3
+38997.9
+39972.5
+41925.0
+43882.0
+45842.9+46
824.8+49774.2+51
743.9+54703.0
38000mass
37884.0
38059.0
560.0
37884
Ian Suydam

45
Charge State Series
700 800 900 1000 1100 1200 1300 1400m/z
+261458.2
+271404.0
+281353.9+29
1307.3
+301263.8
+311223.0
+321184.7
+331148.9
+351083.5
+361053.3
+38997.9
+39972.5
+41925.0
+43882.0
+45842.9+46
824.8+49774.2+51
743.9+54703.0
850 900 950 1000 1050 1100 1150m/z
+331154.3+34
1120.4
+351088.5+36
1058.2
+371029.6
+381002.6
+39976.9+40
952.5+41929.3
+42907.3
+43886.1
+44866.1+45
846.7
+46828.4
38000mass
37884.0
38059.0
560.0
37884
38059
Ian Suydam

46
75 kDa Protein Charge States011105_4139 #281-317 RT: 10.11-11.44 AV: 37 NL: 1.76E6T: + p ESI Full ms [ 550.00-2000.00]
1000 1200 1400 1600 1800 2000m/z
0
10
20
30
40
50
60
70
80
90
100
Rel
ativ
e A
bund
ance
+381967.73
+391917.27
+401869.47
+421780.60
+431739.13
+451662.07
+481558.27
+511466.53
+521438.60
+551360.00
+561335.67
+601246.87
+621206.40
+661133.27
0.5 x 2 mm Michrom Peptide CapTrap

47
75 kD Protein Deconvoluted# 1 RT: 0.00 P: + NL: 2.82E7T: + p ESI Full ms [ 550.00-2000.00]
65000 70000 75000 80000 85000
mass
0
5
10
15
20
25
30
35
40
45
50
55
60
65
70
75
80
85
90
95
100
Rel
ativ
e A
bund
ance
74742 Da

48
High-Resolution MS – Q-Tof
O O
OOO
OMeMe
H
Me Me
H
O
Me
NH2
O
M (neutral)C19H29NO8
MW 399.1893[M+Na]+
C19H29NO8NaMW 422.1791
421 422 423 424m/z0
100
%
031103_12403_AH 132 (2.257) Sm (SG, 2x3.00); Sb (5,40.00 ); 3.29e3422.1802
423.1803422.6927 424.1900
Andrew Hinman

49
Centroided Spectrum
O O
OOO
OMeMe
H
Me Me
H
O
Me
NH2
O
M (neutral)C19H29NO8
MW 399.1893[M+Na]+
C19H29NO8NaMW 422.1791
421 422 423 424m/z0
100
%
0
100
%
031103_12403_AH 132 (2.257) AM (Cen,4, 80.00, Ar,5000.0,0.002.26e4422.1798
423.1862
422.6973 424.1897
031103_12403_AH 132 (2.257) Sm (SG, 2x3.00); Sb (5,40.00 ); 3.29e3422.1802
423.1803422.6927 424.1900
Andrew Hinman

50
Elemental Composition Report
[M+Na]+C19H29NO8NaMW 422.1791
Andrew Hinman
O O
OOO
OMeMe
H
Me Me
H
O
Me
NH2
O

51
Elemental Composition Report
[M+Na]+C19H29NO8NaMW 422.1791
Andrew Hinman
O O
OOO
OMeMe
H
Me Me
H
O
Me
NH2
O
422.1791 amu, 1.7 ppm, C19H29NO8Na

52
ESI-MS of Epothilone C
Epothilone C from E. coli culture broth
[M+H]+ = 478.26[M+Na]+ = 500.24
[M+H]+
[M+Na]+
C28H39NO5SMW: 477.25
OHO
HO N
S
O
O
Chris Boddy

53
Epothilone C Biosynthesis is Reconstituted in E. coli
14C Radio-TLC assay
1 2 31. EpoC standard2. 2 mM substrate3. Negative control
LC/MS analysisExtracted Ion Chromatogram of [M+H]+
epothilone C standard
E. coli with 2 mM substrate
Epothilone C is observed in metabolically engineered E. coli cultures Chris Boddy

54
Characterization by Isotopic Derivatization
12C propionate feeding
12Cm/z = 478.3
13Cm/z = 481.3
SNAC
OH
OO
O
Na
**
* *OHO
HO
O
O
N
SN
S
* = 13C
EIC
Chris Boddy

55
13C Propionate increases mass by 3 Da
13C propionate feeding
SNAC
OH
OO
O
Na
**
* *OHO
HO
O
O
N
SN
S
* = 13C
EIC
Chris Boddy
12Cm/z = 478.3
13Cm/z = 481.3

Conclusion
ResourcesAcknowledgements

57
http://mass-spec.stanford.edu

58
http://mass-spec.stanford.edu

59
Online Tools & Linkshttp://mass-spec.stanford.edu

60
Acknowledgements• Wandless Lab
Hiroko Tanaka • Waymouth Lab
Ned BowdenKuo-Wei Huang
• Stack LabRussell PrattXavi Ribas
• Collman LabRichard Decreau
• Kool LabGreg Miller
• Boxer LabIan Suydam
• DuBois LabAndrew Hinman
• Khosla LabChris Boddy
SUMS:• Andrew Guzzetta• Michael Kitamura
MS Committee:• Pehr Harbury• Peter Jackson• Chaitan Khosla• Al Smith• Tom Wandless
Funding:• Stanford Bio-X Initiative• Vincent & Stella Coates
Foundation