Genetically engineered bacteria: chemical factories of the future?
-
Upload
greg-crowther -
Category
Documents
-
view
1.107 -
download
1
description
Transcript of Genetically engineered bacteria: chemical factories of the future?

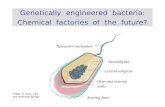
Genetically engineered bacteria:
Chemical factories of the future?
Relocation mechanism
Assembly line
Central computer
Security fence
Outer and internal walls
Image: G. Karp, Cell and molecular biology

Gregory J. Crowther, Ph.D.
Acting Lecturer
Mary E. Lidstrom, Ph.D.
Frank Jungers Professor of Chemical Engineering

The chemical industry today
• supplies chemicals for many manufactured goods
• employs many scientists and engineers
• based on chemicals derived from petroleum
- not a renewable resource- supplied by volatile areas of the world- many produce hazardous wastes
www.hr/tuzla/slike

Possible solution:Use bacteria as chemical factories
Starting materials
Value-added products
• Self-replicating multistage catalysts • Environmentally benign• Use renewable starting materials (feedstocks)

Advantages of bacteria vs. other cells
• Relatively small and simple
• Reproduce quickly
• Tremendous metabolic / catalytic diversity
www.milebymile.com/main/United_States/Wyoming/
- thrive in extreme environments
- use nutrients unavailable to other organisms

Potential products
• Fuels
• Natural products (complex synthesis)
• Engineered products
www.myhealthshack.net; www.acehardware.com
- hydrogen (H2)- methane (CH4)- methanol (CH3OH)- ethanol (CH3CH2OH)
- starting materials for polymers (rubber, plastic, fabrics)- specialty chemicals (chiral)- bulk chemicals (C4 acids)
- vitamins- therapeutic agents- pigments- amino acids- viscosifiers- industrial enzymes- PHAs (biodegradable plastics)

Limitations of naturally occurring bacteria
Bacteria are evolved for survival in competitive natural environments, not for production of chemicals desired by humans!
coolgov.com
- are optimized for low nutrient levels
- have defense systems
- don’t naturally make everything we need

Redesigning bacteria
Goal: optimize industrially valuable parameters.
• Redirect metabolism to specific products
• Remove unwanted products
- storage products
- excretion products
- defense systemspyo.oulu.fi

Metabolic engineering(a form of genetic
engineering)
DNAGene 1 Gene 2 Gene 3
Enzyme 1 Enzyme 2 Enzyme 3A B C D
A
DNA

DNAGene 1 Gene 2 Gene 3
Enzyme 1 Enzyme 2 Enzyme 3A B C D
A
Deleting a gene
DNA

DNAGene 1 Gene 2 Gene 3
Enzyme 1 Enzyme 2 Enzyme 3A B C D
A
Deleting a gene
DNA X

DNAGene 1 Gene 2 Gene 3
Enzyme 1 Enzyme 2 Enzyme 3A B C D
A
Deleting a gene
DNA X
X

DNAGene 1 Gene 2 Gene 3
Enzyme 1 Enzyme 2 Enzyme 3A B C D
A
Deleting a gene
DNA X
X X

Adding a new gene
DNAGene 1 Gene 2 Gene 3
Enzyme 1 Enzyme 2 Enzyme 3A B C D
A
DNA

Adding a new gene
DNAGene 1 Gene 2 Gene 3
Enzyme 1 Enzyme 2 Enzyme 3A B C D
A
Gene 4

Adding a new gene
DNAGene 1 Gene 2 Gene 3
Enzyme 1 Enzyme 2 Enzyme 3A B C D
A
Gene 4
Enzy
me 4

Adding a new gene
DNAGene 1 Gene 2 Gene 3
Enzyme 1 Enzyme 2 Enzyme 3A B C D
A
Gene 4
Enzy
me 4
E

How are genetic changes made?
Most common approach:
1. Put a gene of interest into a stable carrier (vector), a circle of DNA called a plasmid.
2. Put the plasmid into a new cell.
Gene 4
plasmid

How are genetic changes made?
plasmid
Gene 4
Most common approach:
1. Put a gene of interest into a stable carrier (vector), a circle of DNA called a plasmid.
2. Put the plasmid into a new cell.

How are genetic changes made?
plasmid
Gene 4
Most common approach:
1. Put a gene of interest into a stable carrier (vector), a circle of DNA called a plasmid.
2. Put the plasmid into a new cell.

How are genetic changes made?
Gene 4
plasmid
Most common approach:
1. Put a gene of interest into a stable carrier (vector), a circle of DNA called a plasmid.
2. Put the plasmid into a new cell.

How are genetic changes made?
DNAGene 1 Gene 2 Gene 3

How are genetic changes made?
DNAGene 1 Gene 2 Gene 3
Gene 4

How are genetic changes made?
DNAGene 1 Gene 2 Gene 3
Gene 4

How are genetic changes made?
DNAGene 1 Gene 2 Gene 3
Gene 4X X

How are genetic changes made?
DNAGene 1 Gene 2 Gene 3Gene 4

Metabolic engineering mishaps: maximizing ethanol production
PFKethanolglucose
PFK was thought to be the rate-limiting enzyme of ethanol production, so its levels were increased via genetic engineering.
Problem: rates of ethanol production did not increase!

Metabolic engineering mishaps: maximizing PHA production
CH2=H4F
Serine Cycle
CH2=H4MPT
H4MPT
CH3OH
HCHOH4F
CO2
PHA
To maximize PHA production in M. extorquens, one might try to knock out the right-hand pathway.
Problems:
• HCHO builds up and is toxic
• Cells can’t generate enough energy for growth
X

Cellular metabolism is very complicated!
• Lots of molecules
• Highly interconnected
• Mathematical models can help us predict the effects of genetic changes
opbs.okstate.edu/~leach/Bioch5853/

Flux balance analysis
AA B
C
D
E
In a steady state, all concentrations are constant. For each compound, production rate = consumption rate.
To get a solution (flux rate for each step), define an objective function (e.g., production of E) to be maximized.
1010
10
10
0
0
10

Edwards & Palsson (2000)
Reference: PNAS 97: 5528-33, 2000.
Used flux balance analysis to predict how well E. coli cells would grow if various genes were deleted.
The graph at left shows their predictions of how fluxes are rerouted in response to gene deletions.

Edwards & Palsson (2000)
Fraction of normal growth rate
Gene deletions that should not affect growth.
Gene deletions that should slow growth.
Gene deletions that
should stop growth.

Edwards & Palsson (2000)
Predictions of whether various E. coli mutants should be able to grow were compared with experimental data on these mutants.
In 68 of 79 cases (86%), the prediction agreed with the experimental data.

Ethical issues
• Is it OK to tamper with the genes of living organisms?
• What are the possible effects on those organisms?
• What are the possible effects on human health?
• What are the possible effects on the environment?

Summary
• Bacteria have great potential as environmentally friendly chemical “factories.”
• Much additional research will be needed for this potential to be fulfilled.
• Further progress will require knowledge of biology, chemistry, engineering, and mathematics.
www.elsevier.com

More informationabout metabolic engineering
depts.washington.edu/mllab
web.mit.edu/bamel
www.genomatica.com
www.metabolix.com
Lidstrom lab (UW)
Stephanopoulos lab (MIT)
Company founded by Palsson (UCSD)
Well-written background info and examples